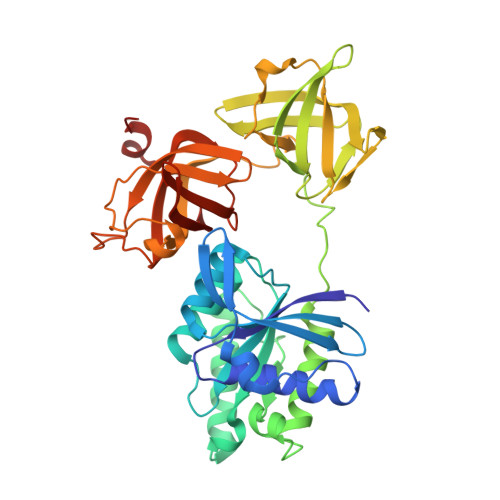
High resolution crystal structure of bovine mitochondrial EF-Tu in complex with GDP.
Andersen, G.R., Thirup, S., Spremulli, L.L., Nyborg, J.(2000) J Mol Biology 297: 421-436
- PubMed: 10715211
- DOI: https://doi.org/10.1006/jmbi.2000.3564
- Primary Citation of Related Structures:
1D2E - PubMed Abstract:
The crystal structure of bovine mitochondrial elongation factor Tu (EF-Tu) in complex with GDP has been determined at a resolution of 1. 94 A. The structure is similar to that of EF-Tu:GDP from Escherichia coli and Thermus aquaticus, but the orientation of the GDP-binding domain 1 is changed relative to domains 2 and 3. Sixteen conserved water molecules common to EF-Tu and other G-proteins in the GDP-binding site are described. These water molecules create a network linking separated parts of the binding pocket. Mitochondrial EF-Tu binds nucleotides less tightly than prokaryotic EF-Tu possibly due to an increased mobility in regions close to the GDP-binding site. The C-terminal extension of mitochondrial EF-Tu has structural similarities with DNA recognising zinc fingers suggesting that the extension may be involved in recognition of RNA.
- Institute of Molecular and Structural Biology, Aarhus University, Gustav Wiedsvej 10C, Aarhus, DK8000, Denmark.
Organizational Affiliation: