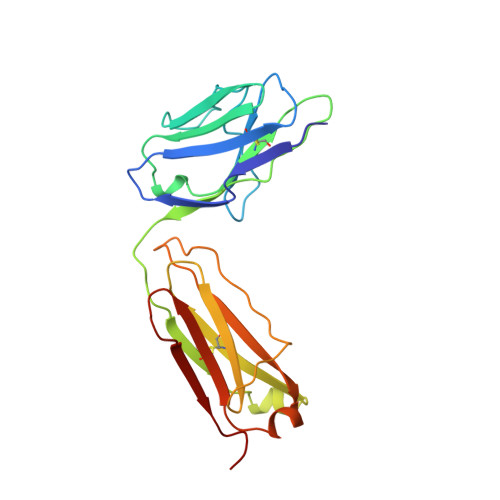
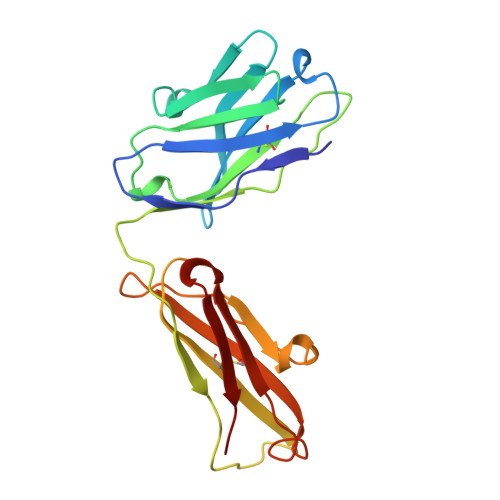
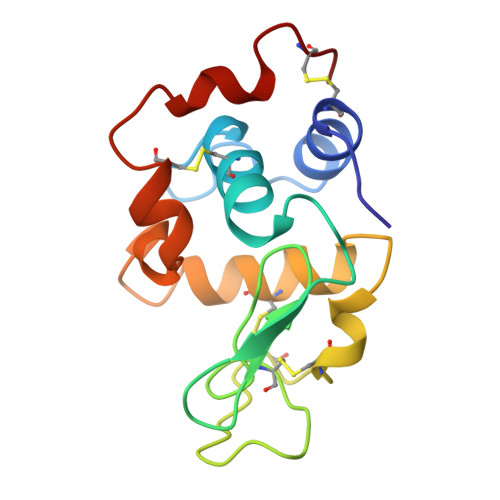
Refined structures of bobwhite quail lysozyme uncomplexed and complexed with the HyHEL-5 Fab fragment.
Chacko, S., Silverton, E.W., Smith-Gill, S.J., Davies, D.R., Shick, K.A., Xavier, K.A., Willson, R.C., Jeffrey, P.D., Chang, C.Y., Sieker, L.C., Sheriff, S.(1996) Proteins 26: 55-65
- PubMed: 8880929 Search on PubMed
- DOI: https://doi.org/10.1002/(SICI)1097-0134(199609)26:1<55::AID-PROT5>3.0.CO;2-F
- Primary Citation Related Structures:
1BQL, 1DKJ, 1DKK - PubMed Abstract:
The HyHEL-5 antibody has more than a thousandfold lower affinity for bobwhite quail lysozyme (BWQL) than for hen egg-white lysozyme (HEL). Four sequence differences exist between BWQL and HEL, of which only one is involved in the interface with the Fab. The structure of bobwhite quail lysozyme has been determined in the uncomplexed state in two different crystal forms and in the complexed state with HyHEL-5, an antihen egg-white lysozyme Fab. Similar backbone conformations are observed in the three molecules of the two crystal forms of uncomplexed BWQL, although they show considerable variability in side-chain conformation. A relatively mobile segment in uncomplexed BWQL is observed to be part of the HyHEL-5 epitope. No major backbone conformational differences are observed in the lysozyme upon complex formation, but side-chain conformational differences are seen in surface residues that are involved in the interface with the antibody. The hydrogen bonding in the interface between BWQL and HyHEL-5 is similar to that in previously determined lysozyme-HyHEL-5 complexes.
- Laboratory of Molecular Biology, NIDDK, NIH, Bethesda, Maryland 20895, USA.
Organizational Affiliation: