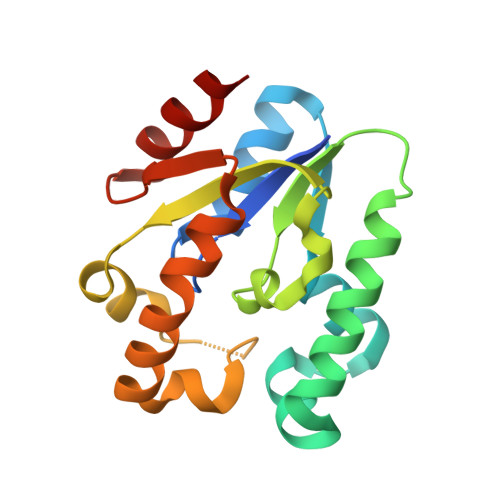
Structural basis for shikimate-binding specificity of Helicobacter pylori shikimate kinase
Cheng, W.C., Chang, Y.N., Wang, W.C.(2005) J Bacteriol 187: 8156-8163
- PubMed: 16291688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.187.23.8156-8163.2005
- Primary Citation Related Structures:
1ZUH, 1ZUI - PubMed Abstract:
Shikimate kinase (EC 2.7.1.71) catalyzes the specific phosphorylation of the 3-hydroxyl group of shikimic acid in the presence of ATP. As the fifth key step in the shikimate pathway for aromatic amino acid biosynthesis in bacteria, fungi, and plants, but not mammals, shikimate kinase represents an attractive target for the development of new antimicrobial agents, herbicides, and antiparasitic agents. Here, we report the 1.8-Angstroms crystal structure of Helicobacter pylori shikimate kinase (HpSK). The crystal structure shows a three-layer alpha/beta fold consisting of a central sheet of five parallel beta-strands flanked by seven alpha-helices. An HpSK-shikimate-PO(4) complex was also determined and refined to 2.3 Angstroms, revealing induced-fit movement from an open to a closed form on substrate binding. Shikimate is located above a short 3(10) helix formed by a strictly conserved motif (GGGXV) after beta(3). Moreover, several highly conserved charged residues including Asp33 (in a conserved DT/SD motif), Arg57, and Arg132 (interacting with shikimate) are identified, guiding the development of novel inhibitors of shikimate kinase.
- Institute of Molecular and Cellular Biology and Department of Life Sciences, National Tsing Hua University, Hsinchu, Taiwan.
Organizational Affiliation: