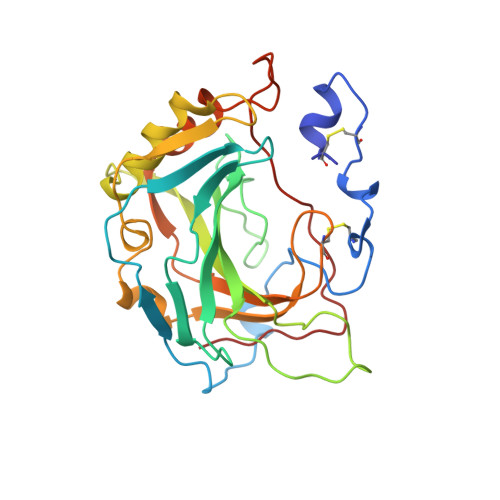
Crystal structure of the secretory form of membrane-associated human carbonic anhydrase IV at 2.8-A resolution.
Stams, T., Nair, S.K., Okuyama, T., Waheed, A., Sly, W.S., Christianson, D.W.(1996) Proc Natl Acad Sci U S A 93: 13589-13594
- PubMed: 8942978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.93.24.13589
- Primary Citation Related Structures:
1ZNC - PubMed Abstract:
It has recently been demonstrated that the C-terminal deletion mutant of recombinant human carbonic anhydrase IV (G267X CA IV) converts the normally glycosylphosphatidylinositol-anchored enzyme into a soluble secretory form which has the same catalytic properties as the membrane-associated enzyme purified from human tissues. We have determined the three-dimensional structure of the secretory form of human CA IV by x-ray crystallographic methods to a resolution of 2.8 A. Although the zinc binding site and the hydrophobic substrate binding pocket of CA IV are generally similar to those of other mammalian isozymes, unique structural differences are found elsewhere in the active site. Two disufide linkages, Cys-6-Cys-11G and Cys-23-Cys-203, stabilize the conformation of the N-terminal domain. The latter disulfide additionally stabilizes an active site loop containing a cis-peptide linkage between Pro-201 and Thr-202 (this loop contains catalytic residue Thr-199). On the opposite side of the active site, the Val-131-Asp-136 segment adopts an extended loop conformation instead of an alpha-helix conformation as found in other isozymes. Finally, the C terminus is surrounded by a substantial electropositive surface potential, which is likely to stabilize the interaction of CA IV with the negatively charged phospholipid headgroups of the membrane. These structural features are unique to CA IV and provide a framework for the design of sulfonamide inhibitors selective for this particular isozyme.
- Department of Chemistry, University of Pennsylvania, Philadelphia 19104-6323, USA.
Organizational Affiliation: