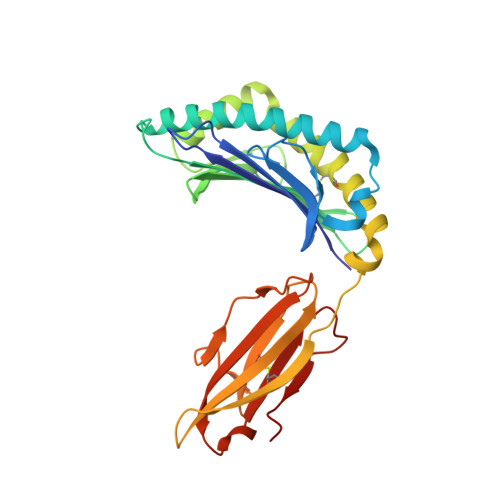
The high resolution structures of highly bulged viral epitopes bound to the major histocompatability class I: Implications for T-cell receptor engagement and T-cell immunodominance
Tynan, F.E., Borg, N.A., Miles, J.J., Beddoe, T., El-Hassen, D., Silins, S.L., van Zuylen, W.J., Purcell, A.W., Kjer-Nielsen, L., McCluskey, J., Burrows, S.R., Rossjohn, J.(2005) J Biological Chem 280: 23900-23909
- PubMed: 15849183 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M503060200
- Primary Citation Related Structures:
1ZHK, 1ZHL - PubMed Abstract:
Although HLA class I alleles can bind epitopes up to 14 amino acids in length, little is known about the immunogenicity or the responding T-cell repertoire against such determinants. Here, we describe an HLA-B*3508-restricted cytotoxic T lymphocyte response to a 13-mer viral epitope (LPEPLPQGQLTAY). The rigid, centrally bulged epitope generated a biased T-cell response. Only the N-terminal face of the peptide bulge was critical for recognition by the dominant clonotype SB27. The SB27 public T-cell receptor (TcR) associated slowly onto the complex between the bulged peptide and the major histocompatibility complex, suggesting significant remodeling upon engagement. The broad antigen-binding cleft of HLA-B*3508 represents a critical feature for engagement of the public TcR, as the narrower binding cleft of HLA-B*3501(LPEPLPQGQLTAY), which differs from HLA-B*3508 by a single amino acid polymorphism (Arg156 --> Leu), interacted poorly with the dominant TcR. Biased TcR usage in this cytotoxic T lymphocyte response appears to reflect a dominant role of the prominent peptide x major histocompatibility complex class I surface.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: