carbonic anhydrase II in complex with ethoxzolamidphenole as sulfonamide inhibitor
Honndorf, V.S., Heine, A., Klebe, G., Supuran, C.T.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Carbonic anhydrase II | 259 | Homo sapiens | Mutation(s): 0 Gene Names: CA2 EC: 4.2.1.1 (PDB Primary Data), 4.2.1.69 (UniProt) | 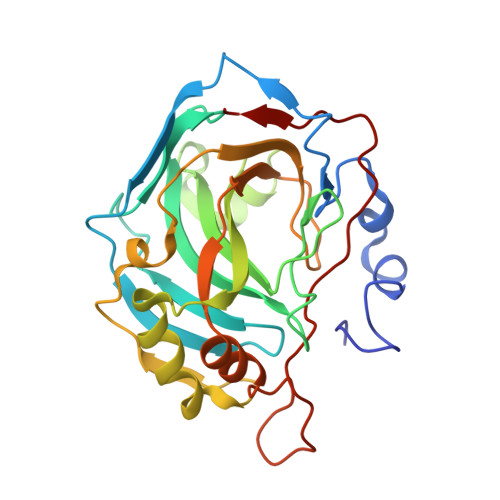 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P00918 (Homo sapiens) Explore P00918 Go to UniProtKB: P00918 | |||||
PHAROS: P00918 GTEx: ENSG00000104267 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00918 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BE7 Query on BE7 | E [auth A] | (4-CARBOXYPHENYL)(CHLORO)MERCURY C7 H5 Cl Hg O2 YFZOUMNUDGGHIW-UHFFFAOYSA-M |  | ||
| ZEC Query on ZEC | C [auth A], D [auth A] | 6-HYDROXY-1,3-BENZOTHIAZOLE-2-SULFONAMIDE C7 H6 N2 O3 S2 NOOBQTYVTDBXTL-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | F [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ZN Query on ZN | B [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 42.39 | α = 90 |
| b = 41.42 | β = 104.54 |
| c = 72.23 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SHELX | model building |
| SHELXL-97 | refinement |
| HKL-2000 | data scaling |
| CNS | phasing |