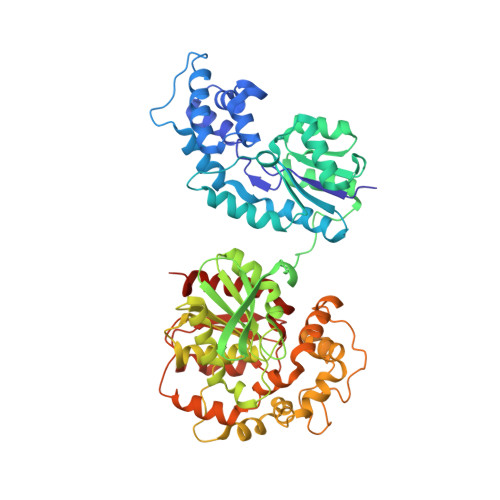
Human soluble epoxide hydrolase: structural basis of inhibition by 4-(3-cyclohexylureido)-carboxylic acids
Gomez, G.A., Morisseau, C., Hammock, B.D., Christianson, D.W.(2006) Protein Sci 15: 58-64
- PubMed: 16322563 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051720206
- Primary Citation Related Structures:
1ZD2, 1ZD3, 1ZD4, 1ZD5 - PubMed Abstract:
X-ray crystal structures of human soluble epoxide hydrolase (sEH) complexed with four different dialkylurea inhibitors bearing pendant carboxylate "tails" of varying length have been determined at 2.3-3.0 A resolution. Similarities among inhibitor binding modes reinforce the proposed roles of Y381 and/or Y465 as general acids that protonate the epoxide ring of the substrate in concert with nucleophilic attack of D333 at the electrophilic epoxide carbon. Additionally, the binding of these inhibitors allows us to model the binding mode of the endogenous substrate 14,15-epoxyeicosatrienoic acid. Contrasts among inhibitor binding modes include opposite orientations of inhibitor binding in the active-site hydrophobic tunnel. Alternative binding orientations observed for this series of inhibitors to human sEH, as well as the binding of certain dialkylurea inhibitors to human sEH and murine sEH, complicate the structure-based design of human sEH inhibitors with potential pharmaceutical applications in the treatment of hypertension. Thus, with regard to the optimization of inhibitor designs targeting human sEH, it is critical that human sEH and not murine sEH be utilized for inhibitor screening, and it is critical that structures of human sEH-inhibitor complexes be determined to verify inhibitor binding orientations that correlate with measured affinities.
- Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: