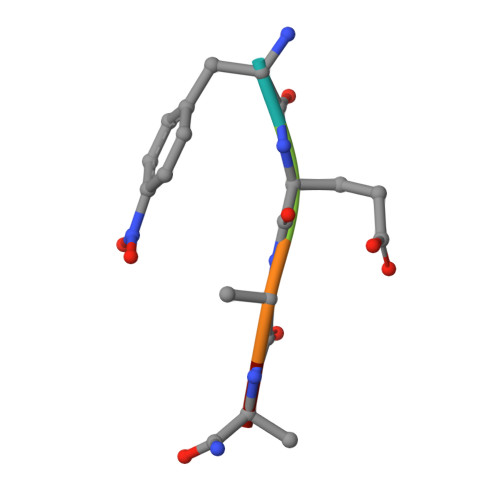
Three-dimensional structures of HIV-1 and SIV protease product complexes.
Rose, R.B., Craik, C.S., Douglas, N.L., Stroud, R.M.(1996) Biochemistry 35: 12933-12944
- PubMed: 8841139 Search on PubMed
- DOI: https://doi.org/10.1021/bi9612733
- Primary Citation Related Structures:
1YTG, 1YTH, 1YTI, 1YTJ - PubMed Abstract:
Strain is eliminated as a factor in hydrolysis of the scissile peptide bond by human immunodeficiency virus (HIV)-1 and simian immunodeficiency virus (SIV), based on the first eight complexes of products of hydrolysis with the enzymes. The carboxyl group generated at the scissile bond interacts with both catalytic aspartic acids. The structures directly suggest the interactions of the gemdiol intermediate with the active site. Based on the structures, the nucleophilic water is displaced stereospecifically by substrate binding toward one catalytic aspartic acid, while the scissile carbonyl becomes hydrogen bonded to the other catalytic aspartic acid in position for hydrolysis. Crystal structures for two N-terminal (P) products and two C-terminal (Q) products provide unambiguous density for the ligands at 2.2-2.6 A resolution and 17-21% R factors. The N-terminal product, Ac-S-L-N-F/, overlaps closely with the N-terminal sequences of peptidomimetic inhibitors bound to the protease. Comparison of the two C-terminal products, /F-L-E-K and /F(NO2)-E-A-Nle-S, indicates that the P2' residue is highly constrained, while the positioning of the P1' and P3' residues are sequence dependent.
- Department of Biochemistry and Biophysics, University of California, San Francisco 94143-0448, USA.
Organizational Affiliation: