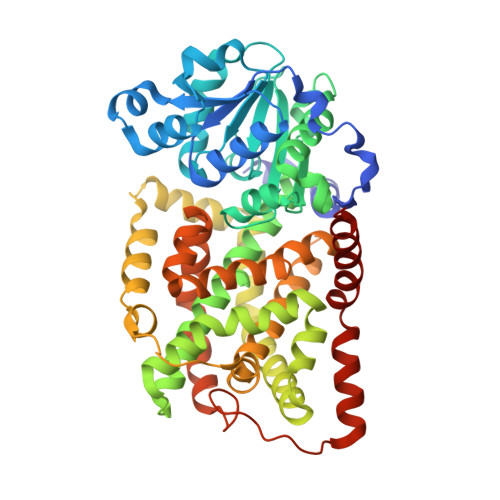
The crystal structure of a bacterial class II ketol-acid reductoisomerase: domain conservation and evolution
Tyagi, R., Duquerroy, S., Navaza, J., Guddat, L.W., Duggleby, R.G.(2005) Protein Sci 14: 3089-3100
- PubMed: 16322583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051791305
- Primary Citation Related Structures:
1YRL - PubMed Abstract:
Ketol-acid reductoisomerase (KARI; EC 1.1.1.86) catalyzes two steps in the biosynthesis of branched-chain amino acids. Amino acid sequence comparisons across species reveal that there are two types of this enzyme: a short form (Class I) found in fungi and most bacteria, and a long form (Class II) typical of plants. Crystal structures of each have been reported previously. However, some bacteria such as Escherichia coli possess a long form, where the amino acid sequence differs appreciably from that found in plants. Here, we report the crystal structure of the E. coli enzyme at 2.6 A resolution, the first three-dimensional structure of any bacterial Class II KARI. The enzyme consists of two domains, one with mixed alpha/beta structure, which is similar to that found in other pyridine nucleotide-dependent dehydrogenases. The second domain is mainly alpha-helical and shows strong evidence of internal duplication. Comparison of the active sites between KARI of E. coli, Pseudomonas aeruginosa, and spinach shows that most residues occupy conserved positions in the active site. E. coli KARI was crystallized as a tetramer, the likely biologically active unit. This contrasts with P. aeruginosa KARI, which forms a dodecamer, and spinach KARI, a dimer. In the E. coli KARI tetramer, a novel subunit-to-subunit interacting surface is formed by a symmetrical pair of bulbous protrusions.
- School of Molecular and Microbial Sciences, The University of Queensland, Brisbane, QLD 4072, Australia.
Organizational Affiliation: