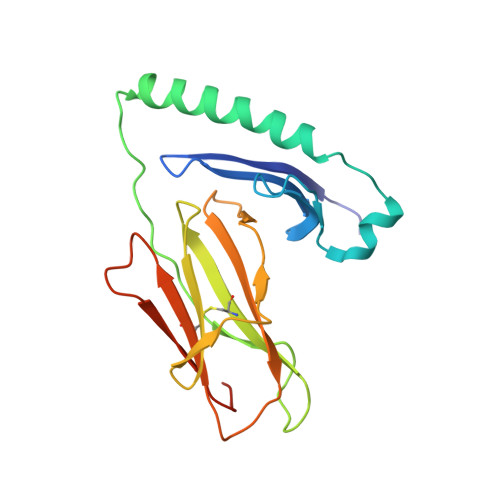
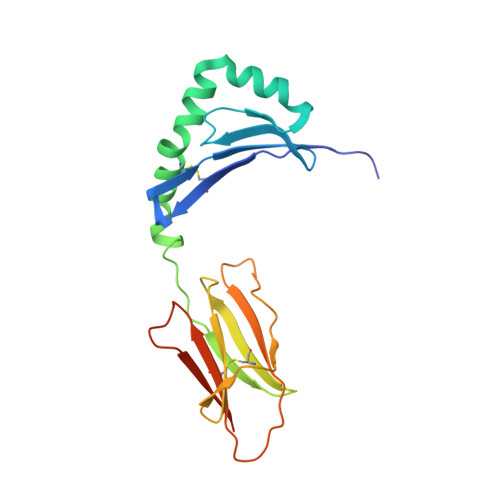
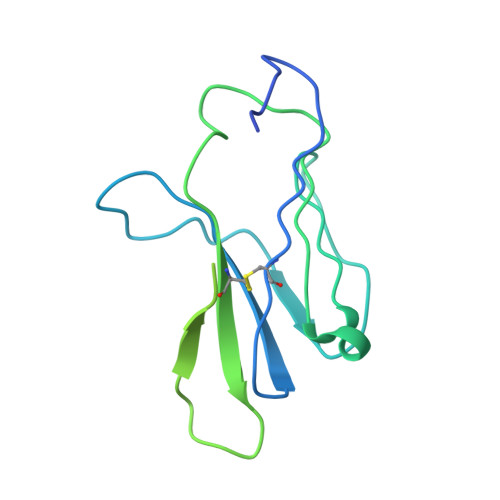
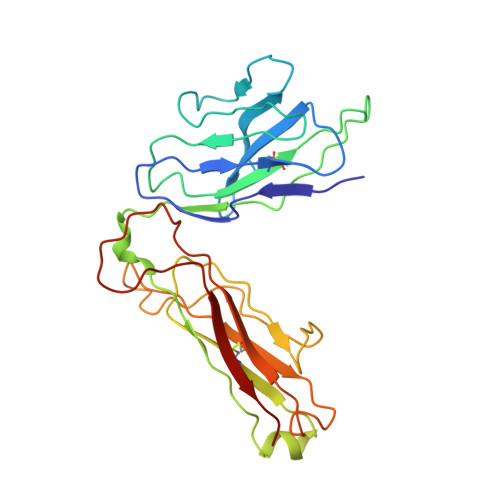
Unconventional topology of self peptide-major histocompatibility complex binding by a human autoimmune T cell receptor.
Hahn, M., Nicholson, M.J., Pyrdol, J., Wucherpfennig, K.W.(2005) Nat Immunol 6: 490-496
- PubMed: 15821740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ni1187
- Primary Citation Related Structures:
1YMM - PubMed Abstract:
Autoimmune diseases are caused by self-reactive lymphocytes that have escaped deletion. Here we have determined the structure of the trimolecular complex for a T cell receptor (TCR) from a patient with multiple sclerosis that causes autoimmunity in transgenic mice. The structure showed a TCR topology notably different from that of antimicrobial TCRs. Rather than being centered on the peptide-major histocompatibility complex, this TCR contacted only the N-terminal peptide segment and made asymmetrical interactions with the major histocompatibility complex helices. The interaction was dominated by the hypervariable complementarity-determining region 3 loops, indicating that unconventional topologies are possible because of the unique complementarity-determining region 3 sequences created during rearrangement. This topology reduces the interaction surface with peptide and alters the geometry for CD4 association. We propose that unusual TCR-binding properties can permit autoreactive T cells to escape deletion.
- Department of Cancer Immunology and AIDS, Dana-Farber Cancer Institute, Boston, Massachusetts 02115, USA.
Organizational Affiliation: