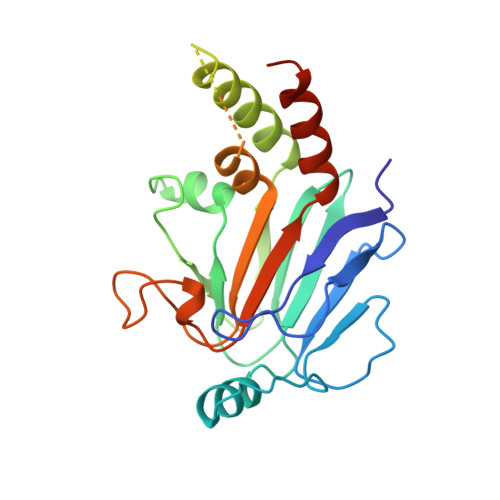
A structural basis for mutational inactivation of the tumour suppressor Smad4.
Shi, Y., Hata, A., Lo, R.S., Massague, J., Pavletich, N.P.(1997) Nature 388: 87-93
- PubMed: 9214508
- DOI: https://doi.org/10.1038/40431
- Primary Citation of Related Structures:
1YGS - PubMed Abstract:
The Smad4/DPC4 tumour suppressor is inactivated in nearly half of pancreatic carcinomas and to a lesser extent in a variety of other cancers. Smad4/DPC4, and the related tumour suppressor Smad2, belong to the SMAD family of proteins that mediate signalling by the TGF-beta/activin/BMP-2/4 cytokine superfamily from receptor Ser/Thr protein kinases at the cell surface to the nucleus. SMAD proteins, which are phosphorylated by the activated receptor, propagate the signal, in part, through homo- and hetero-oligomeric interactions. Smad4/DPC4 plays a central role as it is the shared hetero-oligomerization partner of the other SMADs. The conserved carboxy-terminal domains of SMADs are sufficient for inducing most of the ligand-specific effects, and are the primary targets of tumorigenic inactivation. We now describe the crystal structure of the C-terminal domain (CTD) of the Smad4/DPC4 tumour suppressor, determined at 2.5 A resolution. The structure reveals that the Smad4/DPC4 CTD forms a crystallographic trimer through a conserved protein-protein interface, to which the majority of the tumour-derived missense mutations map. These mutations disrupt homo-oligomerization in vitro and in vivo, indicating that the trimeric assembly of the Smad4/DPC4 CTD is critical for signalling and is disrupted by tumorigenic mutations.
- Cellular Biochemistry and Biophysics Program, Memorial Sloan-Kettering Cancer Center, New York 10021, USA.
Organizational Affiliation: