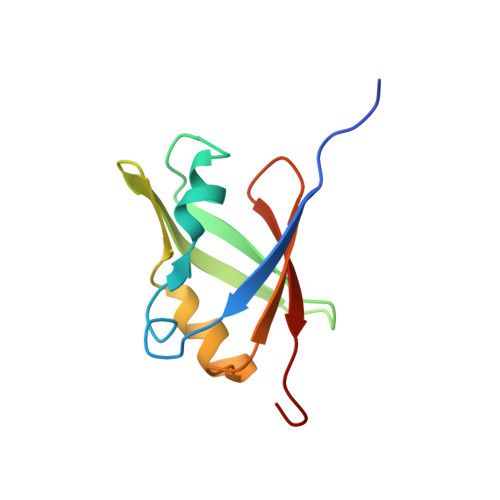
Structure of the B3 domain from Arabidopsis thaliana protein At1g16640
Waltner, J.K., Peterson, F.C., Lytle, B.L., Volkman, B.F.(2005) Protein Sci 14: 2478-2483
- PubMed: 16081658 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051606305
- Primary Citation Related Structures:
1YEL - PubMed Abstract:
A novel DNA binding motif, the B3 domain, has been identified in a number of transcription factors specific to higher plant species, and was recently found to define a new protein fold. Here we report the second structure of a B3 domain, that of the Arabidopsis thaliana protein, At1g16640. As part of an effort to 'rescue' structural genomics targets deemed unsuitable for structure determination as full-length proteins, we applied a combined bioinformatic and experimental strategy to identify an optimal construct containing a predicted conserved domain. By screening a series of N- and C-terminally truncated At1g16640 fragments, we isolated a stable folded domain that met our criteria for structural analysis by NMR spectroscopy. The structure of the B3 domain of At1g16640 consists of a seven-stranded beta-sheet arranged in an open barrel and two short alpha-helices, one at each end of the barrel. While At1g16640 is quite distinct from previously characterized B3 domain proteins in terms of amino acid sequence similarity, it adopts the same novel fold that was recently revealed by the RAV1 B3 domain structure. However, putative DNA-binding elements conserved in B3 domains from the RAV, ARF, and ABI3/VP1 subfamilies are largely absent in At1g16640, perhaps suggesting that B3 domains could function in contexts other than transcriptional regulation.
- Department of Biochemistry and Center for Eukaryotic Structural Genomics, Medical College of Wisconsin, 8701 Watertown Plank Road, Milwaukee, Wisconsin 53226, USA.
Organizational Affiliation: