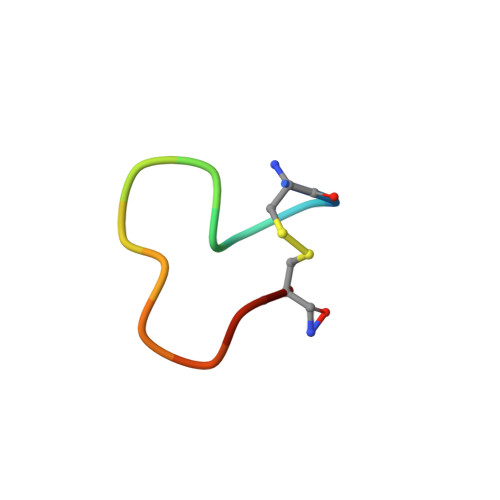
Structural studies and model membrane interactions of two peptides derived from bovine lactoferricin
Nguyen, L.T., Schibli, D.J., Vogel, H.J.(2005) J Pept Sci 11: 379-389
- PubMed: 15635665
- DOI: https://doi.org/10.1002/psc.629
- Primary Citation Related Structures:
1Y58, 1Y5C - PubMed Abstract:
The powerful antimicrobial properties of bovine lactoferricin (LfcinB) make it attractive for the development of new antimicrobial agents. An 11-residue linear peptide portion of LfcinB has been reported to have similar antimicrobial activity to lactoferricin itself, but with lower hemolytic activity. The membrane-binding and membrane-perturbing properties of this peptide were studied together with an amidated synthetic version with an added disulfide bond, which was designed to confer increased stability and possibly activity. The antimicrobial and cytotoxic properties of the peptides were measured against Staphylococcus aureus and Escherichia coli and by hemolysis assays. The peptides were also tested in an anti-cancer assay against neuroblastoma cell lines. Vesicle disruption caused by these LfcinB derivatives was studied using the fluorescent reporter molecule calcein. The extent of burial of the two Trp residues in membrane mimetic environments were quantitated by fluorescence. Finally, the solution NMR structures of the peptides bound to SDS micelles were determined to provide insight into their membrane bound state. The cyclic peptide was found to have greater antimicrobial potency than its linear counterpart. Consistent with this property, the two Trp residues of the modified peptide were suggested to be embedded deeper into the membrane. Although both peptides adopt an amphipathic structure without any regular alpha-helical or beta-sheet conformation, the 3D-structures revealed a clearer partitioning of the cationic and hydrophobic faces for the cyclic peptide.
- Structural Biology Research Group, Department of Biological Sciences, University of Calgary, Calgary, Alberta, T2N 1N4 Canada.
Organizational Affiliation: