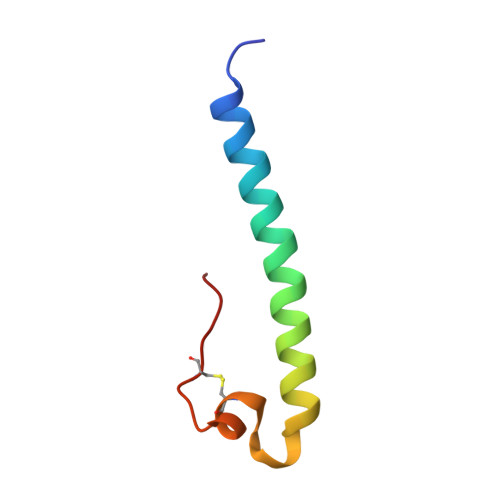
Crystal structure of a pivotal domain of human syncytin-2, a 40 million years old endogenous retrovirus fusogenic envelope gene captured by primates.
Renard, M., Varela, P.F., Letzelter, C., Duquerroy, S., Rey, F.A., Heidmann, T.(2005) J Mol Biology 352: 1029-1034
- PubMed: 16140326 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.058
- Primary Citation Related Structures:
1Y4M - PubMed Abstract:
HERV-FRD is a human endogenous retrovirus that entered the human genome 40 million years ago. Its envelope gene, syncytin-2, was diverted by an ancestral host most probably because of its fusogenic property, for a role in placenta morphogenesis. It was maintained in a functional state in all primate branches as a bona fide cellular gene, submitted to a very low mutation rate as compared to infectious retrovirus genomes. The structure of the syncytin-2 protein thus provides a good insight into that of the oldest mammalian retroviral envelope. Here, we report the crystal structure of a central fragment of its "fossil" ectodomain, allowing a remarkable superposition with the structures of the corresponding domains of present-day infectious retroviruses, in spite of a more than 60% divergent sequence. These results suggest the existence of a unique structural solution selected by these proteins for their fusogenic function.
- Rétrovirus Endogènes et Eléments Rétroïdes des Eucaryotes Supérieurs, UMR 8122 CNRS, Institut Gustave Roussy, 94805 Villejuif, France.
Organizational Affiliation: