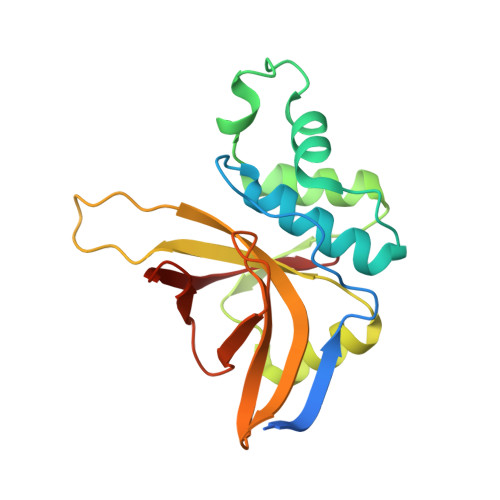
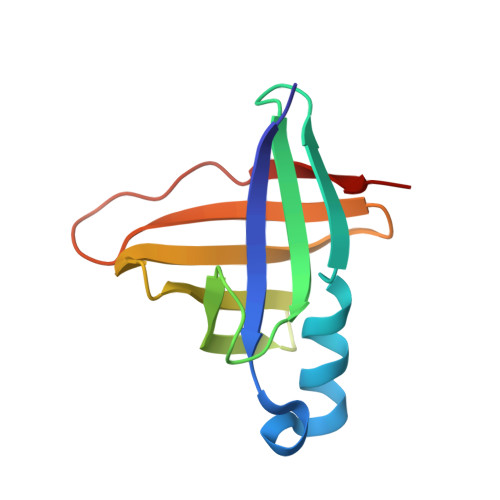
A comparison of staphostatin B with standard mechanism serine protease inhibitors.
Filipek, R., Potempa, J., Bochtler, M.(2005) J Biological Chem 280: 14669-14674
- PubMed: 15644332 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411792200
- Primary Citation Related Structures:
1Y4H - PubMed Abstract:
Staphostatins are the endogenous, highly specific inhibitors of staphopains, the major secreted cysteine proteases from Staphylococcus aureus. We have previously shown that staphostatins A and B are competitive, active site-directed inhibitors that span the active site clefts of their target proteases in the same orientation as substrates. We now report the crystal structure of staphostatin B in complex with wild-type staphopain B at 1.9 A resolution. In the complex structure, the catalytic residues are found in exactly the positions that would be expected for uncomplexed papain-type proteases. There is robust, continuous density for the staphostatin B binding loop and no indication for cleavage of the peptide bond that comes closest to the active site cysteine of staphopain B. The carbonyl carbon atom C of this peptide bond is 4.1 A away from the active site cysteine sulfur Sgamma atom. The carbonyl oxygen atom O of this peptide bond points away from the putative oxyanion hole and lies almost on a line from the Sgamma atom to the C atom. The arrangement is strikingly similar to the "ionmolecule" arrangement for the complex of papain-type enzymes with their substrates but differs significantly from the arrangement conventionally assumed for the Michaelis complex of papain-type enzymes with their substrates and also from the arrangement that is crystallographically observed for complexes of standard mechanism inhibitors and their target serine proteases.
- International Institute of Molecular and Cell Biology, ul. Trojdena 4, 02-109 Warsaw, Poland.
Organizational Affiliation: