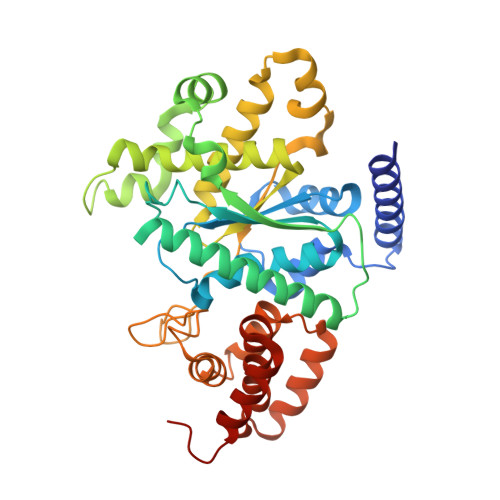
A tyrosyl-tRNA synthetase adapted to function in group I intron splicing by acquiring a new RNA binding surface.
Paukstelis, P.J., Coon, R., Madabusi, L., Nowakowski, J., Monzingo, A., Robertus, J., Lambowitz, A.M.(2005) Mol Cell 17: 417-428
- PubMed: 15694342 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.12.026
- Primary Citation Related Structures:
1Y42 - PubMed Abstract:
We determined a 1.95 A X-ray crystal structure of a C-terminally truncated Neurospora crassa mitochondrial tyrosyl-tRNA synthetase (CYT-18 protein) that functions in splicing group I introns. CYT-18's nucleotide binding fold and intermediate alpha-helical domains superimpose on those of bacterial TyrRSs, except for an N-terminal extension and two small insertions not found in nonsplicing bacterial enzymes. These additions surround the cyt-18-1 mutation site and are sites of suppressor mutations that restore splicing, but not synthetase activity. Highly constrained models based on directed hydroxyl radical cleavage assays show that the group I intron binds at a site formed in part by the three additions on the nucleotide binding fold surface opposite that which binds tRNATyr. Our results show how essential proteins can progressively evolve new functions.
- Institute for Cellular and Molecular Biology, Department of Chemistry and Biochemistry and Section of Molecular Genetics and Microbiology, School of Biological Sciences, University of Texas, Austin, TX 78712, USA.
Organizational Affiliation: