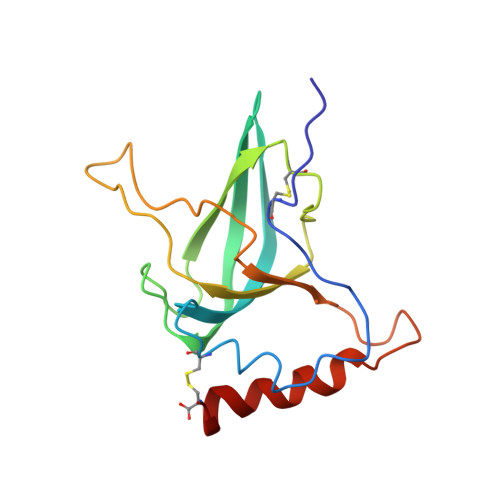
Functional Insights from the Structure of the Multifunctional C345C Domain of C5 of Complement
Bramham, J., Thai, C.-T., Soares, D.C., Uhrin, D., Ogata, R.T., Barlow, P.N.(2005) J Biological Chem 280: 10636-10645
- PubMed: 15598652 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M413126200
- Primary Citation Related Structures:
1XWE - PubMed Abstract:
The complement protein C5 initiates assembly of the membrane attack complex. This remarkable process results in lysis of target cells and is fundamental to mammalian defense against infection. The 150-amino acid residue domain at the C terminus of C5 (C5-C345C) is pivotal to C5 function. It interacts with enzymes that convert C5 to C5b, the first step in the assembly of the membrane attack complex; it also binds to the membrane attack complex components C6 and C7 with high affinity. Here a recombinant version of this C5-C345C domain is shown to adopt the oligosaccharide/oligonucleotide binding fold, with two helices packed against a five-stranded beta-barrel. The structure is compared with those from the netrin-like module family that have a similar fold. Residues critical to the interaction with C5-convertase cluster on a mobile, hydrophobic inter-strand loop that protrudes from the open face of the beta-barrel. The opposite, helix-dominated face of C5-C345C carries a pair of exposed hydrophobic side chains adjacent to a striking negatively charged patch, consistent with affinity for positively charged factor I modules in C6 and C7. Modeling of homologous domains from complement proteins C3 and C4, which do not participate in membrane attack complex assembly, suggests that this provisionally identified C6/C7-interacting face is indeed specific to C5.
- Schools of Chemistry and Biological Sciences, University of Edinburgh, West Mains Road, Edinburgh EH9 3JJ, Scotland, UK.
Organizational Affiliation: