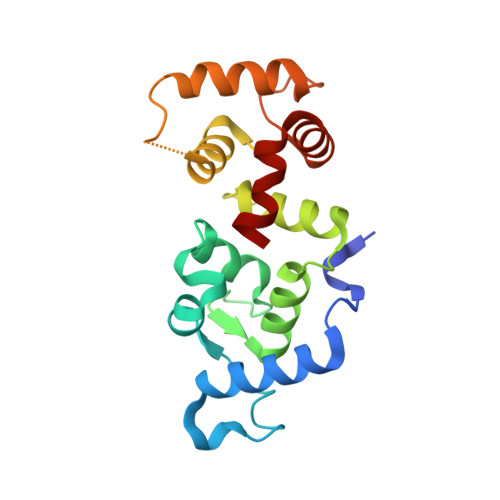
Structural and Biochemical Characterization of CIB1 Delineates a New Family of EF-hand-containing Proteins
Gentry, H.R., Singer, A.U., Betts, L., Yang, C., Ferrara, J.D., Sondek, J., Parise, L.V.(2005) J Biological Chem 280: 8407-8415
- PubMed: 15574431 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411515200
- Primary Citation Related Structures:
1XO5 - PubMed Abstract:
CIB1 (CIB) is an EF-hand-containing protein that binds multiple effector proteins, including the platelet alphaIIbbeta3 integrin and several serine/threonine kinases and potentially modulates their function. The crystal structure for Ca(2+)-bound CIB1 has been determined at 2.0 A resolution and reveals a compact alpha-helical protein containing four EF-hands, the last two of which bind calcium ions in the standard fashion seen in many other EF-hand proteins. CIB1 shares high structural similarity with calcineurin B and the neuronal calcium sensor (NCS) family of EF-hand-containing proteins. Most importantly, like calcineurin B and NCS proteins, which possess a large hydrophobic pocket necessary for ligand binding, CIB1 contains a hydrophobic pocket that has been implicated in ligand binding by previous mutational analysis. However, unlike several NCS proteins, Ca(2+)-bound CIB1 is largely monomeric whether bound to a relevant peptide ligand or ligand-free. Differences in structure, oligomeric state, and phylogeny define a new family of CIB1-related proteins that extends from arthropods to humans.
- Department of Pharmacology, The University of North Carolina, Chapel Hill 27599-7365, USA.
Organizational Affiliation: