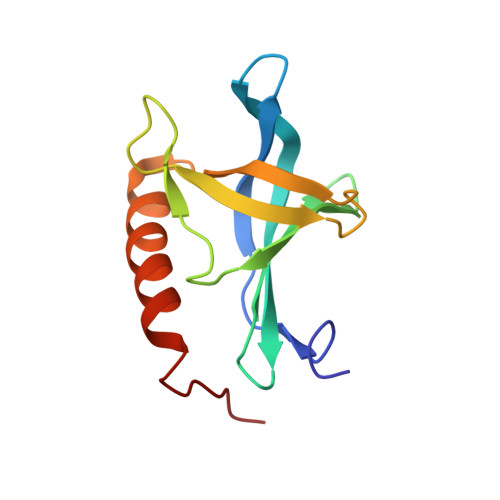
Solution structure of the Ran-binding domain 2 of RanBP2 and its interaction with the C terminus of Ran.
Geyer, J.P., Doker, R., Kremer, W., Zhao, X., Kuhlmann, J., Kalbitzer, H.R.(2005) J Mol Biology 348: 711-725
- PubMed: 15826666 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.02.033
- Primary Citation Related Structures:
1XKE - PubMed Abstract:
The termination of export processes from the nucleus to the cytoplasm in higher eukaryotes is mediated by binding of the small GTPase Ran as part of the export complexes to the Ran-binding domains (RanBD) of Ran-binding protein 2 (RanBP2) of the nuclear pore complex. So far, the structures of the first RanBD of RanBP2 and of RanBP1 in complexes with Ran have been known from X-ray crystallographic studies. Here we report the NMR solution structure of the uncomplexed second RanBD of RanBP2. The structure shows a pleckstrin homology (PH) fold featuring two almost orthogonal beta-sheets consisting of three and four strands and an alpha-helix sitting on top. This is in contrast to the RanBD in the crystal structure complexes in which one beta-strand is missing. That is probably due to the binding of the C-terminal alpha-helix of Ran to the RanBD in these complexes. To analyze the interaction between RanBD2 and the C terminus of Ran, NMR-titration studies with peptides comprising the six or 28 C-terminal residues of Ran were performed. While the six-residue peptide alone does not bind to RanBD2 in a specific manner, the 28-residue peptide, including the entire C-terminal helix of Ran, binds to RanBD2 in a manner analogous to the crystal structures. By solving the solution structure of the 28mer peptide alone, we confirmed that it adopts a stable alpha-helical structure like in native Ran and therefore serves as a valid model of the Ran C terminus. These results support current models that assume recognition of the transport complexes by the RanBDs through the Ran C terminus that is exposed in these complexes.
- Institut für Biophysik und physikalische Biochemie, Universität Regensburg, Universitätsstrasse 31, D-93053 Regensburg, Germany.
Organizational Affiliation: