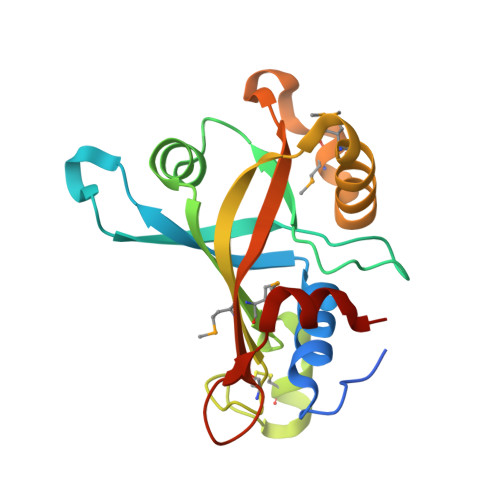
The crystal structure of CREG, a secreted glycoprotein involved in cellular growth and differentiation
Sacher, M., Di Bacco, A., Lunin, V.V., Ye, Z., Wagner, J., Gill, G., Cygler, M.(2005) Proc Natl Acad Sci U S A 102: 18326-18331
- PubMed: 16344469 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0505071102
- Primary Citation Related Structures:
1XHN - PubMed Abstract:
The cellular repressor of E1A-stimulated genes (CREG) is a secreted glycoprotein that inhibits proliferation and enhances differentiation of human embryonal carcinoma cells. CREG binds to the cation-independent mannose 6-phosphate (M6P)/insulin-like growth factor II (IGF2) receptor (IGF2R) (M6P/IGF2R), and this receptor has been shown to be required for CREG-induced growth suppression. To better understand CREG function in cellular growth and differentiation, we solved the 3D crystal structure of this protein to 1.9-A resolution. CREG forms a tight homodimeric complex, and CREG monomers display a beta-barrel fold. The three potential glycosylation sites on CREG map to a confined patch opposite the dimer interface. Thus, dimerization of glycosylated CREG likely presents a bivalent ligand for the M6P/IGF2R. Closely related structural homologs of CREG are FMN-binding split-barrel fold proteins that bind flavin mononucleotide. Our structure shows that the putative flavin mononucleotide-binding pocket in CREG is sterically blocked by a loop and several key bulky residues. A mutant of CREG lacking a part of this loop maintained overall structure and dimerization, as well as M6P/IGF2R binding, but lost the growth suppression activity of WT CREG. Thus, analysis of a structure-based mutant of CREG revealed that binding to M6P/IGF2R, while necessary, is not sufficient for CREG-induced growth suppression. These findings indicate that CREG utilizes a known fold for a previously undescribed function [corrected]
- Montreal Proteomics Network, 740 Doctor Penfield, Montreal, QC, Canada H3A 1A4. michael.sacher@bri.nrc.ca
Organizational Affiliation: