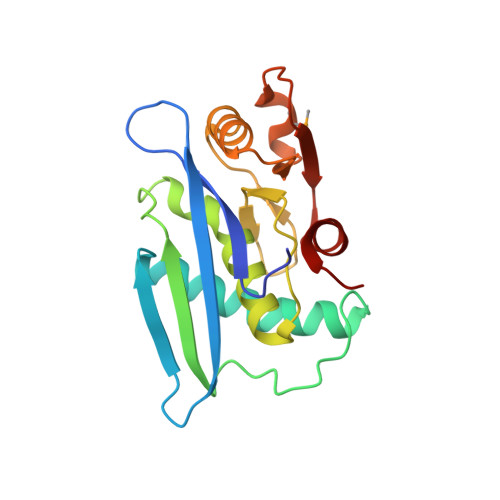
The active site of a lon protease from Methanococcus jannaschii distinctly differs from the canonical catalytic Dyad of Lon proteases.
Im, Y.J., Na, Y., Kang, G.B., Rho, S.H., Kim, M.K., Lee, J.H., Chung, C.H., Eom, S.H.(2004) J Biological Chem 279: 53451-53457
- PubMed: 15456757 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M410437200
- Primary Citation Related Structures:
1XHK - PubMed Abstract:
ATP-dependent Lon proteases catalyze the degradation of various regulatory proteins and abnormal proteins within cells. Methanococcus jannaschii Lon (Mj-Lon) is a homologue of Escherichia coli Lon (Ec-Lon) but has two transmembrane helices within its N-terminal ATPase domain. We solved the crystal structure of the proteolytic domain of Mj-Lon using multiwavelength anomalous dispersion, refining it to 1.9-angstroms resolution. The structure displays an overall fold conserved in the proteolytic domain of Ec-Lon; however, the active site shows uniquely configured catalytic Ser-Lys-Asp residues that are not seen in Ec-Lon, which contains a catalytic dyad. In Mj-Lon, the C-terminal half of the beta4-alpha2 segment is an alpha-helix, whereas it is a beta-strand in Ec-Lon. Consequently, the configurations of the active sites differ due to the formation of a salt bridge between Asp-547 and Lys-593 in Mj-Lon. Moreover, unlike Ec-Lon, Mj-Lon has a buried cavity in the region of the active site containing three water molecules, one of which is hydrogen-bonded to catalytic Ser-550. The geometry and environment of the active site residues in Mj-Lon suggest that the charged Lys-593 assists in lowering the pK(a) of the Ser-550 hydroxyl group via its electrostatic potential, and the water in the cavity acts as a proton acceptor during catalysis. Extensive sequence alignment and comparison of the structures of the proteolytic domains clearly indicate that Lon proteases can be classified into two groups depending on active site configuration and the presence of DGPSA or (D/E)GDSA consensus sequences, as represented by Ec-Lon and Mj-Lon.
- Department of Life Science, Gwangju Institute of Science and Technology, Gwangju 500-712, Korea.
Organizational Affiliation: