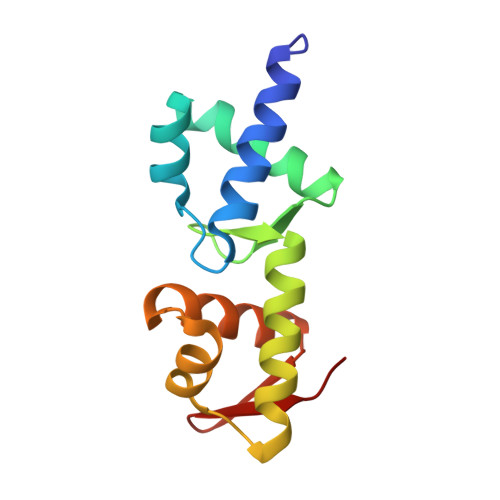
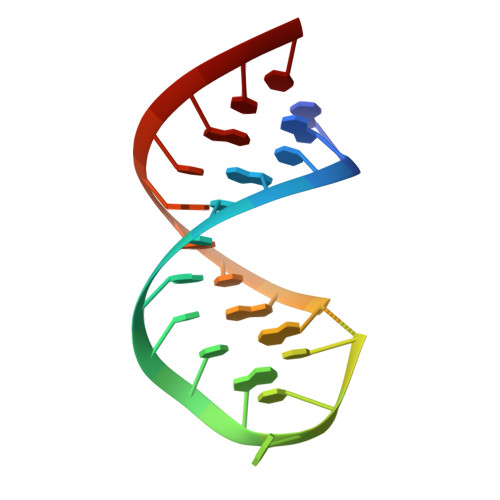
Structural basis for mRNA recognition by elongation factor SelB
Yoshizawa, S., Rasubala, L., Ose, T., Kohda, D., Fourmy, D., Maenaka, K.(2005) Nat Struct Mol Biol 12: 198-203
- PubMed: 15665870 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb890
- Primary Citation Related Structures:
1WSU - PubMed Abstract:
In bacteria, incorporation of selenocysteine, the 21(st) amino acid, into proteins requires elongation factor SelB, which has the unusual property of binding to both transfer RNA (tRNA) and mRNA. SelB binds to an mRNA hairpin formed by the selenocysteine insertion sequence (SECIS) with extremely high specificity, the molecular basis of which has been unknown. We have determined the crystal structure of the mRNA-binding domain of SelB in complex with SECIS RNA at a resolution of 2.3 A. This is the first example of a complex between an RNA and a winged-helix (WH) domain, a motif found in many DNA-binding proteins and recently discovered in RNA-binding proteins. Notably, RNA binding does not induce a major conformational change in the WH motif. The structure reveals a new mode of RNA recognition with a geometry that allows the complex to wrap around the small ribosomal subunit.
- Laboratoire de RMN, Institut de Chimie des Substances Naturelles, Centre National de la Recherche Scientifique, 1 Avenue de la Terrasse, 91190 Gif-sur-Yvette, France. yoshizawa@icsn.cnrs-gif.fr
Organizational Affiliation: