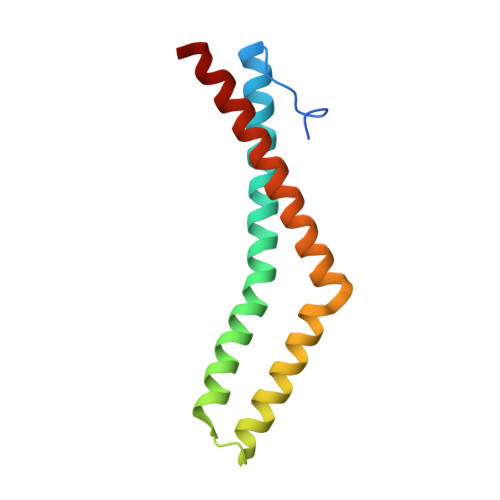
Structure of the conserved cytoplasmic C-terminal domain of occludin: identification of the ZO-1 binding surface.
Li, Y., Fanning, A.S., Anderson, J.M., Lavie, A.(2005) J Mol Biology 352: 151-164
- PubMed: 16081103 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.017
- Primary Citation Related Structures:
1WPA, 1XAW - PubMed Abstract:
Occludin is a transmembrane protein localized at tight junctions whose functions are complex yet poorly understood. Current evidence supports a role for occludin in both the formation of the paracellular barrier and in cell signaling. While the N-terminal extracellular domains of occludin mediate homotypic adhesion, the distal C-terminal cytoplasmic domain of occludin controls protein targeting and endocytosis. The C terminus can also bind to the scaffolding proteins ZO-1, ZO-2, ZO-3, cingulin, the membrane trafficking protein VAP33, and the cytoskeletal protein F-actin, suggesting an important role for this domain. This domain is highly homologous to an important functional domain in the C terminus of the ELL family of RNA polymerase II transcription factors. To explore the function of occludin, we determined the high-resolution crystal structure of its C-terminal distal cytoplasmic domain. The structure comprises three helices that form two separate anti-parallel coiled-coils and a loop that packs tightly against one of the coiled-coils. Using in vitro binding studies and site-directed mutagenesis, we have identified a large positively charged surface that contains the binding site for ZO-1, and this surface is required for proper localization of occludin to cell-cell junctions. On the basis of sequence conservation, we predict that occludin domains from different species and the C-terminal domain of the ELL transcription factors share a very similar structure. Our results provide a model to further test the function of occludin and its binding to other proteins.
- Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, Chicago, IL 60607, USA.
Organizational Affiliation: