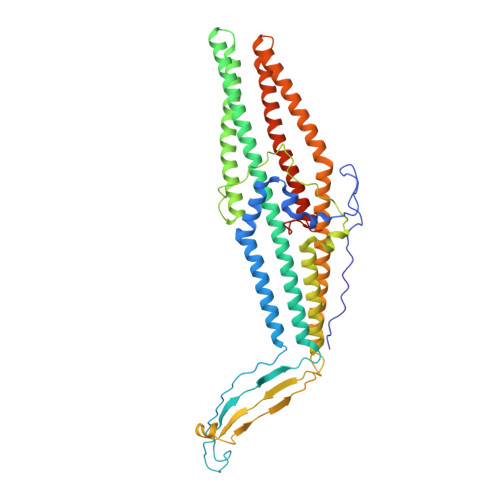
Crystal structure of the drug discharge outer membrane protein, OprM, of Pseudomonas aeruginosa: dual modes of membrane anchoring and occluded cavity end
Akama, H., Kanemaki, M., Yoshimura, M., Tsukihara, T., Kashiwagi, T., Yoneyama, H., Narita, S., Nakagawa, A., Nakae, T.(2004) J Biological Chem 279: 52816-52819
- PubMed: 15507433 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C400445200
- Primary Citation Related Structures:
1WP1 - PubMed Abstract:
The OprM lipoprotein of Pseudomonas aeruginosa is a member of the MexAB-OprM xenobiotic-antibiotic transporter subunits that is assumed to serve as the drug discharge duct across the outer membrane. The channel structure must differ from that of the porin-type open pore because the protein facilitates the exit of antibiotics but not the entry. For better understanding of the structure-function linkage of this important pump subunit, we studied the x-ray crystallographic structure of OprM at the 2.56-angstroms resolution. The overall structure exhibited trimeric assembly of the OprM monomer that consisted mainly of two domains: the membrane-anchoring beta-barrel and the cavity-forming alpha-barrel. OprM anchors the outer membrane by two modes of membrane insertions. One is via the covalently attached NH(2)-terminal fatty acids and the other is the beta-barrel structure consensus on the outer membrane-spanning proteins. The beta-barrel had a pore opening with a diameter of about 6-8 angstroms, which is not large enough to accommodate the exit of any antibiotics. The periplasmic alpha-barrel was about 100 angstroms long formed mainly by a bundle of alpha-helices that formed a solvent-filled cavity of about 25,000 angstroms(3). The proximal end of the cavity was tightly sealed, thereby not permitting the entry of any molecule. The result of this structure was that the resting state of OprM had a small outer membrane pore and a tightly closed periplasmic end, which sounds plausible because the protein should not allow free access of antibiotics. However, these observations raised another unsolved problem about the mechanism of opening of the OprM cavity ends. The crystal structure offers possible mechanisms of pore opening and pump assembly.
- Department of Molecular Life Science, Tokai University School of Medicine, Isehara 259-1193, Japan.
Organizational Affiliation: