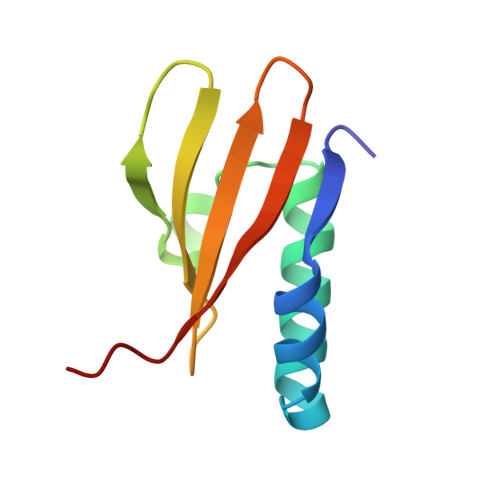
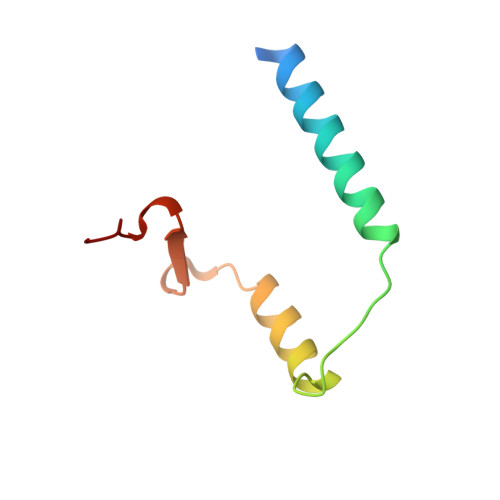
Crystal structure of archaeal toxin-antitoxin RelE-RelB complex with implications for toxin activity and antitoxin effects
Takagi, H., Kakuta, Y., Okada, T., Yao, M., Tanaka, I., Kimura, M.(2005) Nat Struct Mol Biol 12: 327-331
- PubMed: 15768033 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb911
- Primary Citation Related Structures:
1WMI - PubMed Abstract:
The Escherichia coli chromosome encodes toxin-antitoxin pairs. The toxin RelE cleaves mRNA positioned at the A-site in ribosomes, whereas the antitoxin RelB relieves the effect of RelE. The hyperthermophilic archaeon Pyrococcus horikoshii OT3 has the archaeal homologs aRelE and aRelB. Here we report the crystal structure of aRelE in complex with aRelB determined at a resolution of 2.3 A. aRelE folds into an alpha/beta structure, whereas aRelB lacks a distinct hydrophobic core and extensively wraps around the molecular surface of aRelE. Neither component shows structural homology to known ribonucleases or their inhibitors. Site-directed mutagenesis suggests that Arg85, in the C-terminal region, is strongly involved in the functional activity of aRelE, whereas Arg40, Leu48, Arg58 and Arg65 play a modest role in the toxin's activity.
- Laboratory of Biochemistry, Department of Bioscience and Biotechnology, Faculty of Agriculture, Graduate School, Kyushu University, Fukuoka 812-8581, Japan.
Organizational Affiliation: