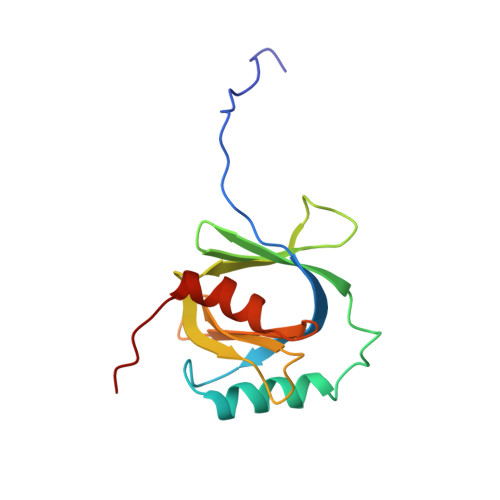
Structure of the C-terminal phosphotyrosine interaction domain of Fe65L1 complexed with the cytoplasmic tail of amyloid precursor protein reveals a novel peptide binding mode
Li, H., Koshiba, S., Hayashi, F., Tochio, N., Tomizawa, T., Kasai, T., Yabuki, T., Motoda, Y., Harada, T., Watanabe, S., Inoue, M., Hayashizaki, Y., Tanaka, A., Kigawa, T., Yokoyama, S.(2008) J Biological Chem 283: 27165-27178
- PubMed: 18650440 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M803892200
- Primary Citation Related Structures:
1WGU, 2ROZ, 2YSZ, 2YT0, 2YT1 - PubMed Abstract:
Fe65L1, a member of the Fe65 family, is an adaptor protein that interacts with the cytoplasmic domain of Alzheimer amyloid precursor protein (APP) through its C-terminal phosphotyrosine interaction/phosphotyrosine binding (PID/PTB) domain. In the present study, the solution structures of the C-terminal PID domain of mouse Fe65L1, alone and in complex with a 32-mer peptide (DAAVTPEERHLSKMQQNGYENPTYKFFEQMQN) derived from the cytoplasmic domain of APP, were determined using NMR spectroscopy. The C-terminal PID domain of Fe65L1 alone exhibits a canonical PID/PTB fold, whereas the complex structure reveals a novel mode of peptide binding. In the complex structure, the NPTY motif forms a type-I beta-turn, and the residues immediately N-terminal to the NPTY motif form an antiparallel beta-sheet with the beta5 strand of the PID domain, the binding mode typically observed in the PID/PTB.peptide complex. On the other hand, the N-terminal region of the peptide forms a 2.5-turn alpha-helix and interacts extensively with the C-terminal alpha-helix and the peripheral regions of the PID domain, representing a novel mode of peptide binding that has not been reported previously for the PID/PTB.peptide complex. The indispensability of the N-terminal region of the peptide for the high affinity of the PID-peptide interaction is consistent with NMR titration and isothermal calorimetry data. The extensive binding features of the PID domain of Fe65L1 with the cytoplasmic domain of APP provide a framework for further understanding of the function, trafficking, and processing of APP modulated by adapter proteins.
- Systems and Structural Biology Center, RIKEN Yokohama Institute, 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: