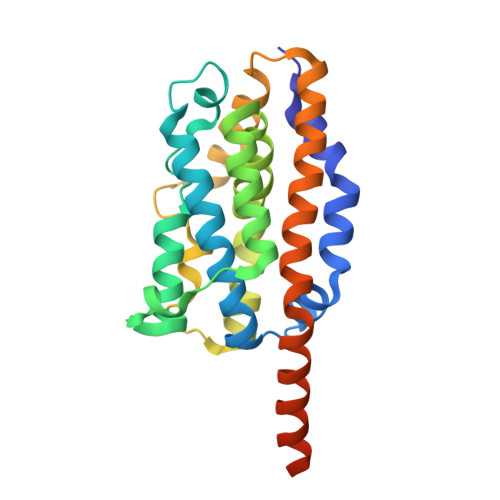
Crystal structure of heme oxygenase-1 from cyanobacterium Synechocystis sp. PCC 6803 in complex with heme
Sugishima, M., Migita, C.T., Zhang, X., Yoshida, T., Fukuyama, K.(2004) Eur J Biochem 271: 4517-4525
- PubMed: 15560792 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.2004.04411.x
- Primary Citation Related Structures:
1WE1 - PubMed Abstract:
Heme oxygenase (HO) catalyzes the oxidative degradation of heme utilizing molecular oxygen and reducing equivalents. In photosynthetic organisms, HO functions in the biosynthesis of such open-chain tetrapyrroles as phyto-chromobilin and phycobilins, which are involved in the signal transduction for light responses and light harvesting for photosynthesis, respectively. We have determined the first crystal structure of a HO-1 from a photosynthetic organism, Synechocystis sp. PCC 6803 (Syn HO-1), in complex with heme at 2.5 A resolution. Heme-Syn HO-1 shares a common folding with other heme-HOs. Although the heme pocket of heme-Syn HO-1 is, for the most part, similar to that of mammalian HO-1, they differ in such features as the flexibility of the distal helix and hydrophobicity. In addition, 2-propanol derived from the crystallization solution occupied the hydrophobic cavity, which is proposed to be a CO trapping site in rat HO-1 that suppresses product inhibition. Although Syn HO-1 and mammalian HO-1 are similar in overall structure and amino acid sequence (57% similarity vs. human HO-1), their molecular surfaces differ in charge distribution. The surfaces of the heme binding sides are both positively charged, but this patch of Syn HO-1 is narrow compared to that of mammalian HO-1. This feature is suited to the selective binding of ferredoxin, the physiological redox partner of Syn HO-1; the molecular size of ferredoxin is approximately 10 kDa whereas the size of NADPH-cytochrome P450 reductase, a reducing partner of mammalian HO-1, is approximately 77 kDa. A docking model of heme-Syn HO-1 and ferredoxin suggests indirect electron transfer from an iron-sulfur cluster in ferredoxin to the heme iron of heme-Syn HO-1.
- Department of Biology, Graduate School of Science, Osaka University, Toyonaka, Osaka, Japan.
Organizational Affiliation: