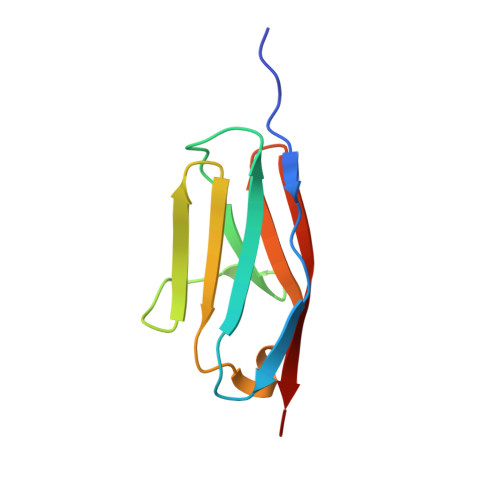
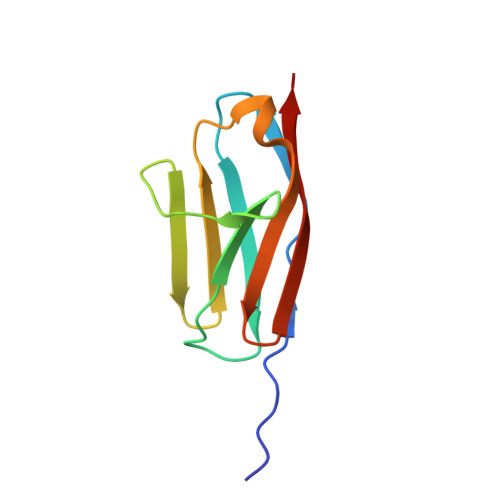
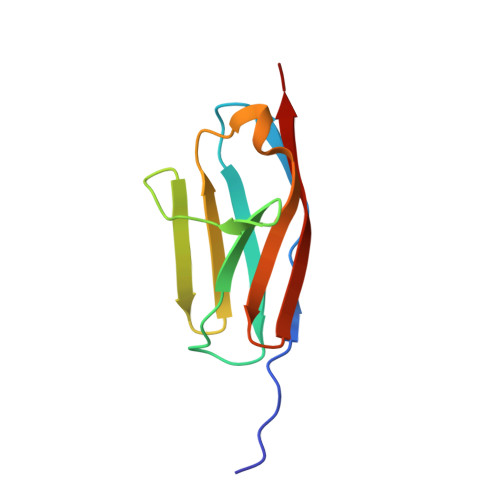
Mechanical Network in Titin Immunoglobulin from Force Distribution Analysis.
Stacklies, W., Vega, M.C., Wilmanns, M., Grater, F.(2009) PLoS Comput Biol 5: 00306
- PubMed: 19282960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pcbi.1000306
- Primary Citation Related Structures:
1WAA - PubMed Abstract:
The role of mechanical force in cellular processes is increasingly revealed by single molecule experiments and simulations of force-induced transitions in proteins. How the applied force propagates within proteins determines their mechanical behavior yet remains largely unknown. We present a new method based on molecular dynamics simulations to disclose the distribution of strain in protein structures, here for the newly determined high-resolution crystal structure of I27, a titin immunoglobulin (IG) domain. We obtain a sparse, spatially connected, and highly anisotropic mechanical network. This allows us to detect load-bearing motifs composed of interstrand hydrogen bonds and hydrophobic core interactions, including parts distal to the site to which force was applied. The role of the force distribution pattern for mechanical stability is tested by in silico unfolding of I27 mutants. We then compare the observed force pattern to the sparse network of coevolved residues found in this family. We find a remarkable overlap, suggesting the force distribution to reflect constraints for the evolutionary design of mechanical resistance in the IG family. The force distribution analysis provides a molecular interpretation of coevolution and opens the road to the study of the mechanism of signal propagation in proteins in general.
- CAS-MPG Partner Institute for Computational Biology, Shanghai, People's Republic of China.
Organizational Affiliation: