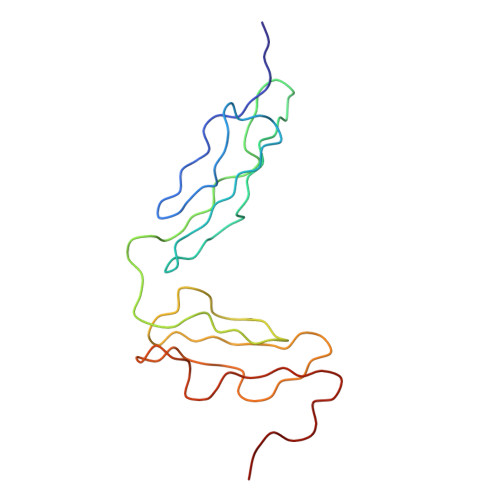
Solution Structure of the Complex between Cr2 Scr 1-2 and C3D of Human Complement: An X-Ray Scattering and Sedimentation Modelling Study.
Gilbert, H.E., Eaton, J.T., Hannan, J.P., Holers, V.M., Perkins, S.J.(2005) J Mol Biology 346: 859
- PubMed: 15713468 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.12.006
- Primary Citation Related Structures:
1W2R, 1W2S - PubMed Abstract:
Complement receptor type 2 (CR2, CD21) forms a tight complex with C3d, a fragment of C3, the major complement component. Previous crystal structures of the C3d-CR2 SCR 1-2 complex and free CR2 SCR 1-2 showed that the two SCR domains of CR2 form contact with each other in a closed V-shaped structure. SCR 1 and SCR 2 are connected by an unusually long eight-residue linker peptide. Medium-resolution solution structures for CR2 SCR 1-2, C3d, and their complex were determined by X-ray scattering and analytical ultracentrifugation. CR2 SCR 1-2 is monomeric. For CR2 SCR 1-2, its radius of gyration R(G) of 2.12(+/-0.05) nm, its maximum length of 10nm and its sedimentation coefficient s20,w(o) of 1.40(+/-0.03) S do not agree with those calculated from the crystal structures, and instead suggest an open structure. Computer modelling of the CR2 SCR1-2 solution structure was based on the structural randomisation of the eight-residue linker peptide joining SCR 1 and SCR 2 to give 9950 trial models. Comparisons with the X-ray scattering curve indicated that the most favoured arrangements for the two SCR domains corresponded to an open V-shaped structure with no contacts between the SCR domains. For C3d, X-ray scattering and sedimentation velocity experiments showed that it exists as a monomer-dimer equilibrium with a dissociation constant of 40 microM. The X-ray scattering curve for monomeric C3d gave an R(G) value of 1.95 nm, and this together with its s20,w(o) value of 3.17 S gave good agreement with the monomeric C3d crystal structure. Modelling of the C3d dimer gave good agreements with its scattering and ultracentrifugation parameters. For the complex, scattering and ultracentrifugation experiments showed that there was no dimerisation, indicating that the C3d dimerisation site was located close to the CR2 SCR 1-2 binding site. The R(G) value of 2.44(+/-0.1) nm, its length of 9 nm and its s20,w(o) value of 3.45(+/-0.01) S showed that its structure was not much more elongated than that of C3d. Calculations with 9950 models of CR2 SCR 1-2 bound to C3d through SCR 2 showed that SCR 1 formed an open V-shaped structure with SCR 2 and was capable of interacting with the surface of C3d. We conclude that the open V-shaped structures formed by CR2 SCR 1-2, both when free and when bound to C3d, are optimal for the formation of a tight two-domain interaction with its ligand C3d.
- Department of Biochemistry and Molecular Biology, Darwin Building, University College London, Gower Street, London WC1E 6BT, UK.
Organizational Affiliation: