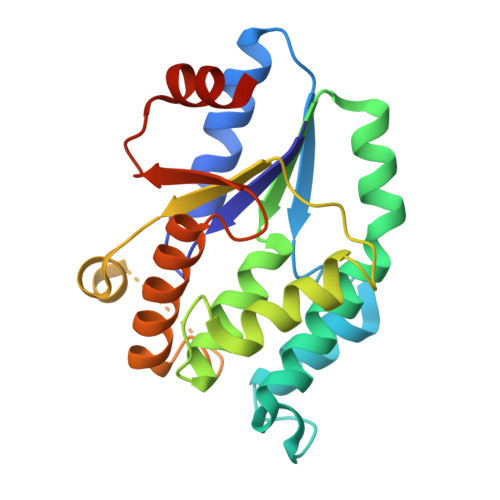
The Crystal Structure of Mycobacterium Tuberculosis Thymidylate Kinase in Complex with 3'-Azidodeoxythymidine Monophosphate Suggests a Mechanism for Competitive Inhibition
Fioravanti, E., Adam, V., Munier-Lehmann, H., Bourgeois, D.(2005) Biochemistry 44: 130
- PubMed: 15628853 Search on PubMed
- DOI: https://doi.org/10.1021/bi0484163
- Primary Citation Related Structures:
1W2G, 1W2H - PubMed Abstract:
Tuberculosis (TB) is the primary cause of mortality among infectious diseases. Mycobacterium tuberculosis thymidylate kinase (TMPK(Mtub)) catalyzes the ATP-dependent phosphorylation of deoxythymidine 5'-monophosphate (dTMP). Essential to DNA replication, this enzyme represents a promising target for developing new drugs against TB, because the configuration of its active site is unique within the TMPK family. Indeed, it has been proposed that, as opposed to other TMPKs, catalysis by TMPK(Mtub) necessitates the transient binding of a magnesium ion coordinating the phosphate acceptor. Moreover, 3'-azidodeoxythymidine monophosphate (AZTMP) is a competitive inhibitor of TMPK(Mtub), whereas it is a substrate for human and other TMPKs. Here, the crystal structures of TMPK(Mtub) in complex with deoxythymidine (dT) and AZTMP were determined to 2.1 and 2.0 A resolution, respectively, and suggest a mechanism for inhibition. The azido group of AZTMP perturbs the induced-fit mechanism normally adopted by the enzyme. Magnesium is prevented from binding, and the resulting electrostatic environment precludes phosphoryl transfer from occurring. Our data provide a model for drug development against tuberculosis.
- LCCP, UMR 5075, IBS-CEA/CNRS/UJF, 41 avenue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: