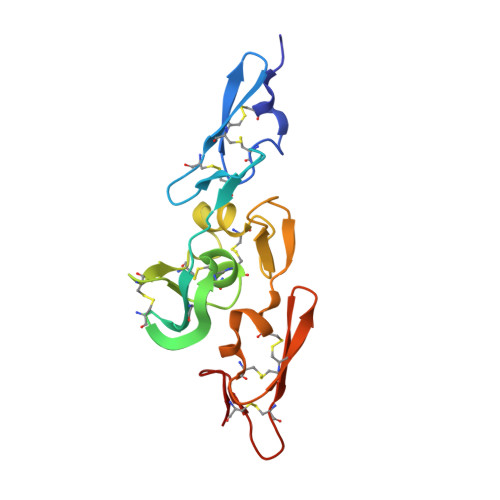
Structure of the Integrin Binding Fragment from Fibrillin-1 Gives New Insights Into Microfibril Organization
Lee, S.S.J., Knott, V., Jovanovi, J., Harlos, K., Grimes, J., Choulier, L., Mardon, H., Stuart, D.I., Handford, P.A.(2004) Structure 12: 717
- PubMed: 15062093 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2004.02.023
- Primary Citation Related Structures:
1UZJ, 1UZK, 1UZP, 1UZQ - PubMed Abstract:
Human fibrillin-1, the major structural protein of extracellular matrix (ECM) 10-12 nm microfibrils, is dominated by 43 calcium binding epidermal growth factor-like (cbEGF) and 7 transforming growth factor beta binding protein-like (TB) domains. Crystal structures reveal the integrin binding cbEGF22-TB4-cbEGF23 fragment of human fibrillin-1 to be a Ca(2+)-rigidified tetragonal pyramid. We suggest that other cbEGF-TB pairs within the fibrillins may adopt a similar orientation to cbEGF22-TB4. In addition, we have located a flexible RGD integrin binding loop within TB4. Modeling, cell attachment and spreading assays, immunocytochemistry, and surface plasmon resonance indicate that cbEGF22 bound to TB4 is a requirement for integrin activation and provide insight into the molecular basis of the fibrillin-1 interaction with alphaVbeta3. In light of our data, we propose a novel model for the assembly of the fibrillin microfibril and a mechanism to explain its extensibility.
- The Henry Wellcome Building of Genomic Medicine, Roosevelt Drive, Oxford OX3 7BN, United Kingdom.
Organizational Affiliation: