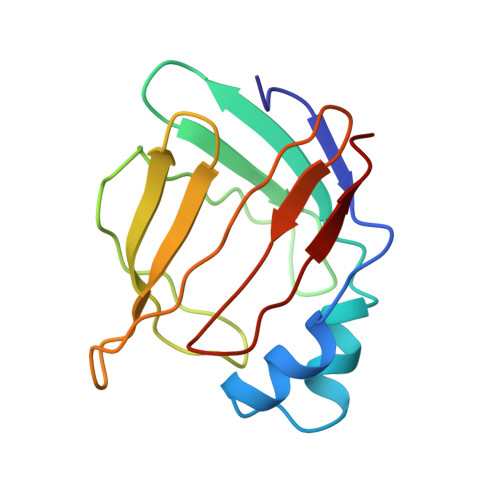
The NMR solution structure of intestinal fatty acid-binding protein complexed with palmitate: application of a novel distance geometry algorithm.
Hodsdon, M.E., Ponder, J.W., Cistola, D.P.(1996) J Mol Biology 264: 585-602
- PubMed: 8969307 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0663
- Primary Citation Related Structures:
1URE - PubMed Abstract:
The three-dimensional solution structure of rat intestinal fatty acid-binding protein (I-FABP) complexed with palmitate has been determined using multidimensional triple-resonance NMR methods. The structure is based on 3889 conformational restraints derived mostly from 3-D 13C- and 15N-resolved nuclear Overhauser (NOESY) experiments. The 3-D NOESY data for this 15.4 kDa complex contained an average of nine possible interpretations per cross-peak. To circumvent this ambiguity, an eight-stage iterative procedure was employed to gradually interpret and introduce unambiguous distance restraints during subsequent rounds of structure calculations. The first stage of this procedure relied critically upon an initial structural model based on the consensus 1H/13C chemical shift-derived secondary structure and a set of symmetry-checked restraints derived from the 3-D 13C-resolved NOESY spectrum. The structures were calculated using DISTGEOM, a program that implements a novel distance geometry algorithm with pairwise Gaussian metrization. A central feature of this algorithm is the use of an iteratively optimized Gaussian distribution for the selection of trial distances, which overcomes the tendency of metrization to produce crushed structures. In addition, this algorithm randomly selects pairwise elements of the distance matrix, which results in an improved sampling of conformational space for a given computational effort. The final family of 20 distance geometry/simulated annealing structures exhibited an average pairwise C(alpha) root-mean-square deviation of 0.98 A, and their stereochemical quality, as assessed by PROCHECK, was comparable to that of 2.5 A X-ray crystal structures. The NMR structure was compared with the X-ray crystal structure of the same ligand/protein complex and was found to be essentially identical within the precision of the results. The NMR structure was also compared with that of the palmitate complex with bovine heart FABP, which shares 30% sequence identity with rat I-FABP. The overall folds were the same, but differences were noted with respect to the presence or absence of apparent conformational heterogeneity and the location and conformation of the bound fatty acid.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110, USA.
Organizational Affiliation: