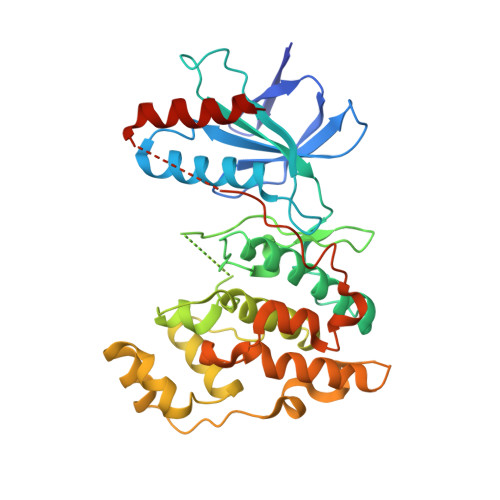
Structural basis for the selective inhibition of JNK1 by the scaffolding protein JIP1 and SP600125
Heo, Y.-S., Kim, S.K., Seo, C.I., Kim, Y.K., Sung, B.-J., Lee, H.S., Lee, J.I., Park, S.-Y., Kim, J.H., Hwang, K.Y., Hyun, Y.-L., Jeon, Y.H., Ro, S., Cho, J.M., Lee, T.G., Yang, C.-H.(2004) EMBO J 23: 2185-2195
- PubMed: 15141161 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600212
- Primary Citation Related Structures:
1UKH, 1UKI - PubMed Abstract:
The c-jun N-terminal kinase (JNK) signaling pathway is regulated by JNK-interacting protein-1 (JIP1), which is a scaffolding protein assembling the components of the JNK cascade. Overexpression of JIP1 deactivates the JNK pathway selectively by cytoplasmic retention of JNK and thereby inhibits gene expression mediated by JNK, which occurs in the nucleus. Here, we report the crystal structure of human JNK1 complexed with pepJIP1, the peptide fragment of JIP1, revealing its selectivity for JNK1 over other MAPKs and the allosteric inhibition mechanism. The van der Waals contacts by the three residues (Pro157, Leu160, and Leu162) of pepJIP1 and the hydrogen bonding between Glu329 of JNK1 and Arg156 of pepJIP1 are critical for the selective binding. Binding of the peptide also induces a hinge motion between the N- and C-terminal domains of JNK1 and distorts the ATP-binding cleft, reducing the affinity of the kinase for ATP. In addition, we also determined the ternary complex structure of pepJIP1-bound JNK1 complexed with SP600125, an ATP-competitive inhibitor of JNK, providing the basis for the JNK specificity of the compound.
- The Division of Drug Discovery, CrystalGenomics, Inc., Daeduk Biocommunity, Jeonmin-dong, Daejon, Korea.
Organizational Affiliation: