Insights into the Phosphoregulation of beta-Secretase Sorting Signal by the VHS Domain of GGA1
Shiba, T., Kametaka, S., Kawasaki, M., Shibata, M., Waguri, S., Uchiyama, Y., Wakatsuki, S.(2004) Traffic 5: 437-448
- PubMed: 15117318 Search on PubMed
- DOI: https://doi.org/10.1111/j.1600-0854.2004.00188.x
- Primary Citation Related Structures:
1UJJ, 1UJK - PubMed Abstract:
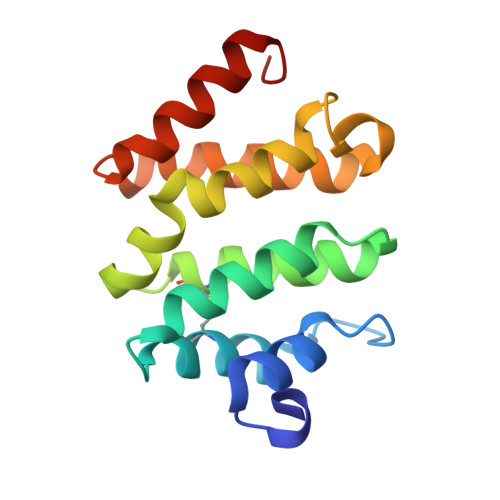
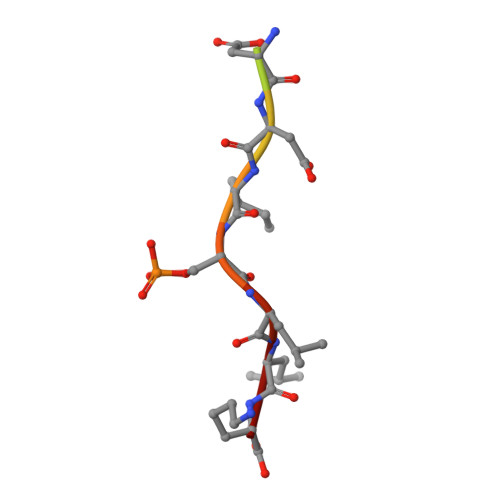
BACE (beta-site amyloid precursor protein cleaving enzyme, beta-secretase) is a type-I membrane protein which functions as an aspartic protease in the production of beta-amyloid peptide, a causative agent of Alzheimer's disease. Its cytoplasmic tail has a characteristic acidic-cluster dileucine motif recognized by the VHS domain of adaptor proteins, GGAs (Golgi-localizing, gamma-adaptin ear homology domain, ARF-interacting). Here we show that BACE is colocalized with GGAs in the trans-Golgi network and peripheral structures, and phosphorylation of a serine residue in the cytoplasmic tail enhances interaction with the VHS domain of GGA1 by about threefold. The X-ray crystal structure of the complex between the GGA1-VHS domain and the BACE C-terminal peptide illustrates a similar recognition mechanism as mannose 6-phosphate receptors except that a glutamine residue closes in to fill the gap created by the shorter BACE peptide. The serine and lysine of the BACE peptide point their side chains towards the solvent. However, phosphorylation of the serine affects the lysine side chain and the peptide backbone, resulting in one additional hydrogen bond and a stronger electrostatic interaction with the VHS domain, hence the reversible increase in affinity.
- Structural Biology Research Center, Photon Factory, Institute of Materials Structure Science, High Energy Accelerator Research Organization (KEK), Tsukuba, Ibaraki 305-0801, Japan.
Organizational Affiliation: