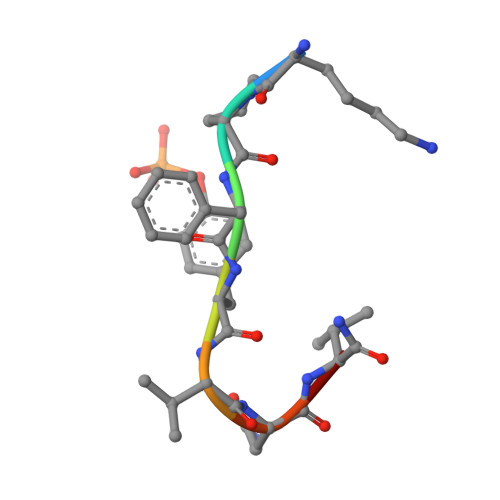
Structural basis for specificity of Grb2-SH2 revealed by a novel ligand binding mode.
Rahuel, J., Gay, B., Erdmann, D., Strauss, A., Garcia-Echeverria, C., Furet, P., Caravatti, G., Fretz, H., Schoepfer, J., Grutter, M.G.(1996) Nat Struct Biol 3: 586-589
- PubMed: 8673601 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0796-586
- Primary Citation Related Structures:
1TZE