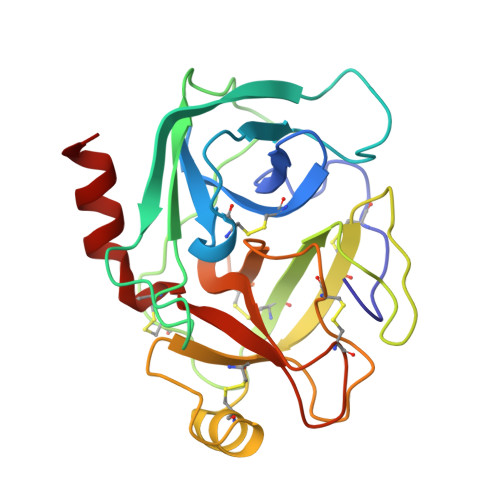
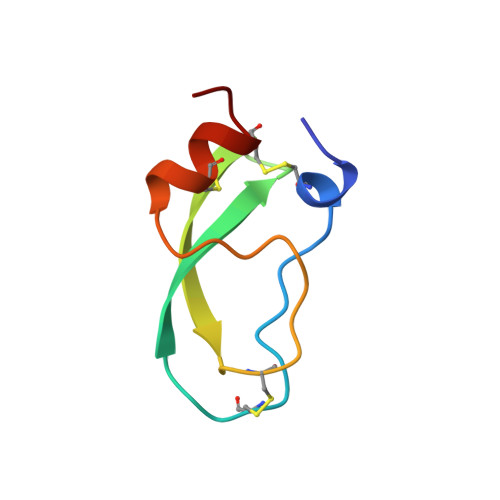
The second Kunitz domain of human tissue factor pathway inhibitor: cloning, structure determination and interaction with factor Xa.
Burgering, M.J., Orbons, L.P., van der Doelen, A., Mulders, J., Theunissen, H.J., Grootenhuis, P.D., Bode, W., Huber, R., Stubbs, M.T.(1997) J Mol Biology 269: 395-407
- PubMed: 9199408 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1029
- Primary Citation Related Structures:
1ADZ, 1TFX - PubMed Abstract:
Tissue Factor Pathway Inhibitor (TFPI) is a 36 kDa glycoprotein that helps maintain haemostasis by inhibiting Factor Xa and the Factor VIIa/Tissue Factor (TF) complex. TFPI contains three tandemly linked Kunitz inhibitor domains, of which the second inhibits factor Xa. We have undertaken a multidisciplinary approach to study the structure and function of the second Kunitz domain of TFPI, with a view towards the rational design of factor Xa inhibitors. Amino acid residues 93 to 154 of the mature TFPI protein, corresponding to the second Kunitz domain (TFPI-kII), were expressed in Escherichia coli. The protein was purified to near homogeneity by ion exchange, hydrophobic interaction, and size exclusion chromatography, respectively. TFPI-kII is a potent factor Xa inhibitor with a Ki of 1.5 x 10(-10) M, a value that does not differ significantly from that of intact TFPI. The three-dimensional structure of TFPI-kII in aqueous solution was determined by 1H nuclear magnetic resonance spectroscopy (NMR). A set of 30 conformers was calculated with the program DIANA using 906 distance constraints derived from nuclear Overhauser effects and 23 dihedral angle constraints. This set, representing the solution structure of TFPI-kII, has an average root-mean-square deviation of 0.78 A for the backbone atoms and 1.38 A for all heavy atoms of residues 1 to 58. The structure of TFPI-kII has also been determined in complex with porcine trypsin using X-ray crystallographic techniques. The complex has been solved to a resolution of 2.6 A, with a final R-factor of 16.2%. Comparison of the NMR derived structure with that of TFPI-kII in complex with trypsin reveals little divergence of the two structures, with the exception of residue Tyr17. Superposition of the trypsin:TFPI-kII complex on factor Xa provides insights into macromolecular determinants for the inhibition of factor Xa. Complexation would require a degree of reorganisation of factor Xa residues, in particular of TyrF99, but also perhaps of the F148-loop. The interaction was further investigated using restrained molecular dynamics. Electrostatic interactions would appear to play a major role. The reorganisation of factor Xa is in contrast to the proposed factor Xa:TAP interaction, where TAP would bind to the "ground state" structure of factor Xa.
- Scientific Development Group, N.V. Organon, Oss, The Netherlands.
Organizational Affiliation: