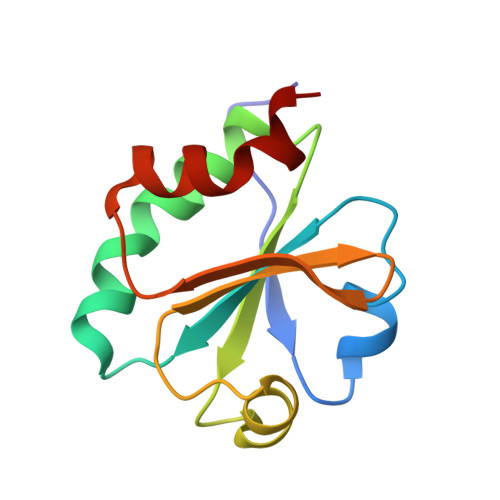
Expression, purification and X-ray crystallographic analysis of thioredoxin from Streptomyces coelicolor.
Stefankova, P., Maderova, J., Barak, I., Kollarova, M., Otwinowski, Z.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 164-168
- PubMed: 16510983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309104032993
- Primary Citation Related Structures:
1T00 - PubMed Abstract:
Thioredoxins are ubiquitous proteins that serve as reducing agents and general protein disulfide reductases. In turn, they are reduced by electrons obtained from the NADPH-containing thioredoxin reductase. Thioredoxins have been isolated and characterized from a large number of organisms. The Gram-positive bacterium Streptomyces coelicolor contains three thioredoxins that are involved in unknown biological processes. trxA from S. coelicolor was cloned and expressed in Escherichia coli and the protein purified and crystallized using the hanging-drop method of vapour diffusion. The crystal structure of thioredoxin A has been determined at 1.5 A resolution using a synchrotron-radiation source. The protein reveals a thioredoxin-like fold with a typical CXXC active site. The crystal exhibits the symmetry of space group P2(1)2(1)2, with unit-cell parameters a = 43.6, b = 71.8, c = 33.2 A.
- Department of Biochemistry, Faculty of Natural Sciences, Comenius University, Mlynska dolina CH-1, 842 15 Bratislava, Slovak Republic.
Organizational Affiliation: