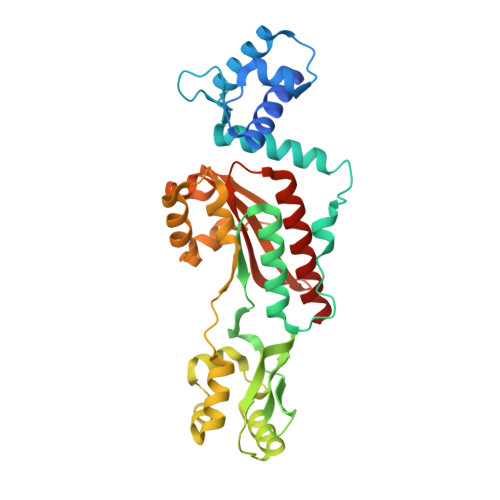
Crystal structure of a heat-inducible transcriptional repressor HrcA from Thermotoga maritima: structural insight into DNA binding and dimerization.
Liu, J., Huang, C., Shin, D.H., Yokota, H., Jancarik, J., Kim, J.S., Adams, P.D., Kim, R., Kim, S.H.(2005) J Mol Biology 350: 987-996
- PubMed: 15979091 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.04.021
- Primary Citation Related Structures:
1STZ - PubMed Abstract:
All cells have a defense mechanism against a sudden heat-shock stress. Commonly, they express a set of proteins that protect cellular proteins from being denatured by heat. Among them, GroE and DnaK chaperones are representative defending systems, and their transcription is regulated by a heat-shock repressor protein HrcA. HrcA repressor controls the transcription of groE and dnaK operons by binding the palindromic CIRCE element, presumably as a dimer, and the activity of HrcA repressor is modulated by GroE chaperones. Here, we report the first crystal structure of a heat-inducible transcriptional repressor, HrcA, from Thermotoga maritima at 2.2A resolution. The Tm_HrcA protein crystallizes as a dimer. The monomer is composed of three domains: an N-terminal winged helix-turn-helix domain (WH), a GAF-like domain, and an inserted dimerizing domain (IDD). The IDD shows a unique structural fold with an anti-parallel beta-sheet composed of three beta-strands sided by four alpha-helices. The Tm_HrcA dimer structure is formed through hydrophobic contact between the IDDs and a limited contact that involves conserved residues between the GAF-like domains. In the overall dimer structure, the two WH domains are exposed, but the conformation of these two domains seems to be incompatible with DNA binding. We suggest that our structure may represent an inactive form of the HrcA repressor. Structural implication on how the inactive form of HrcA may be converted to the active form by GroEL binding to a conserved C-terminal sequence region of HrcA is discussed.
- Berkeley Structural Genomics Center, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: