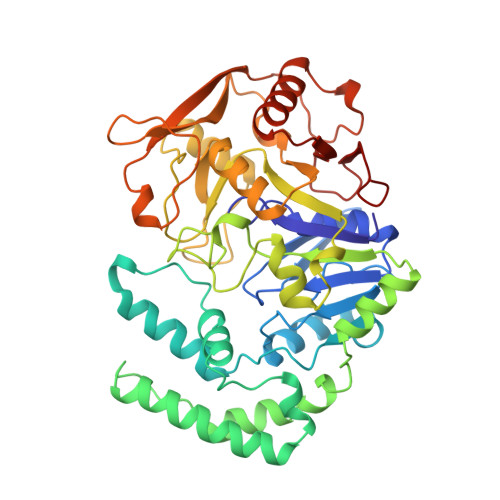
The mode of action and the structure of a herbicide in complex with its target: binding of activated hydantocidin to the feedback regulation site of adenylosuccinate synthetase.
Fonne-Pfister, R., Chemla, P., Ward, E., Girardet, M., Kreuz, K.E., Honzatko, R.B., Fromm, H.J., Schar, H.P., Grutter, M.G., Cowan-Jacob, S.W.(1996) Proc Natl Acad Sci U S A 93: 9431-9436
- PubMed: 8790347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.93.18.9431
- Primary Citation Related Structures:
1SON, 1SOO - PubMed Abstract:
(+)-Hydantocidin, a recently discovered natural spironucleoside with potent herbicidal activity, is shown to be a proherbicide that, after phosphorylation at the 5' position, inhibits adenylosuccinate synthetase, an enzyme involved in de novo purine synthesis. The mode of binding of hydantocidin 5'-monophosphate to the target enzyme was analyzed by determining the crystal structure of the enzyme-inhibitor complex at 2.6-A resolution. It was found that adenylosuccinate synthetase binds the phosphorylated compound in the same fashion as it does adenosine 5'-monophosphate, the natural feedback regulator of this enzyme. This work provides the first crystal structure of a herbicide-target complex reported to date.
- Pharmaceutical Division, Ciba-Geigy Ltd., Basel, Switzerland.
Organizational Affiliation: