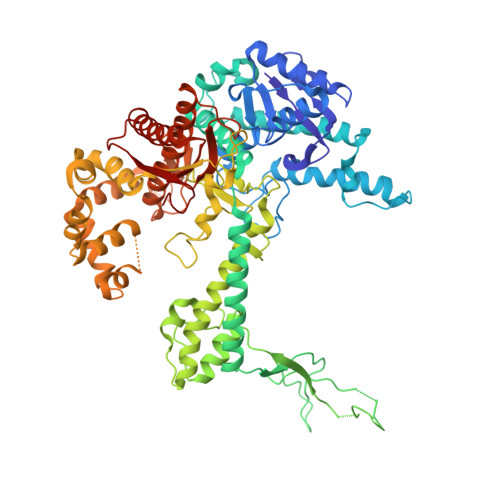
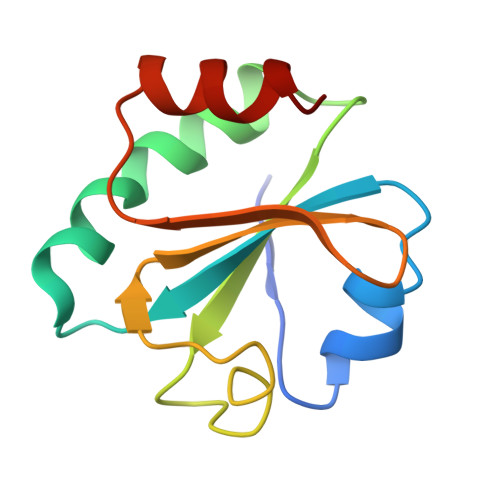
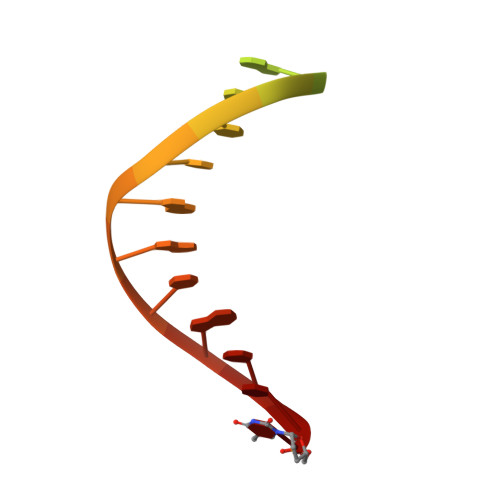
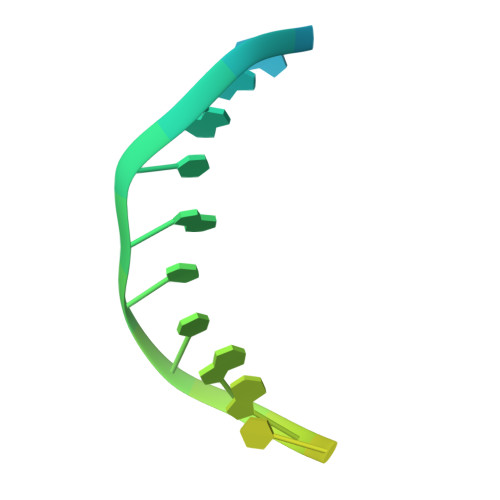
Nucleotide insertion opposite a cis-syn thymine dimer by a replicative DNA polymerase from bacteriophage T7.
Li, Y., Dutta, S., Doublie, S., Bdour, H.M., Taylor, J.S., Ellenberger, T.(2004) Nat Struct Mol Biol 11: 784-790
- PubMed: 15235589 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb792
- Primary Citation Related Structures:
1SKR, 1SKS, 1SKW, 1SL0, 1SL1, 1SL2 - PubMed Abstract:
Ultraviolet-induced DNA damage poses a lethal block to replication. To understand the structural basis for this, we determined crystal structures of a replicative DNA polymerase from bacteriophage T7 in complex with nucleotide substrates and a DNA template containing a cis-syn cyclobutane pyrimidine dimer (CPD). When the 3' thymine is the templating base, the CPD is rotated out of the polymerase active site and the fingers subdomain adopts an open orientation. When the 5' thymine is the templating base, the CPD lies within the polymerase active site where it base-pairs with the incoming nucleotide and the 3' base of the primer, while the fingers are in a closed conformation. These structures reveal the basis for the strong block of DNA replication that is caused by this photolesion.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, 240 Longwood Avenue, Boston, Massachusetts 02115, USA.
Organizational Affiliation: