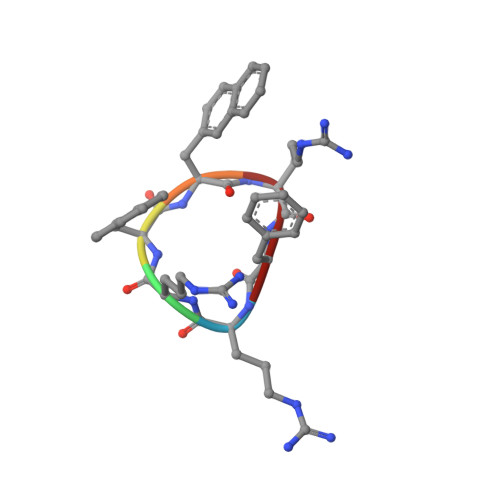
Structure of the antimicrobial, cationic hexapeptide cyclo(RRWWRF) and its analogues in solution and bound to detergent micelles.
Appelt, C., Wessolowski, A., Soderhall, J.A., Dathe, M., Schmieder, P.(2005) Chembiochem 6: 1654-1662
- PubMed: 16075425 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.200500095
- Primary Citation Related Structures:
1QVK, 1QVL, 1SKI, 1SKK, 1SKL - PubMed Abstract:
Antimicrobial, cationic peptides are abundant throughout nature as part of many organisms' defence against microorganisms. They exhibit a large variety of sequences and structural motifs and are thought to act by rupturing the bacterial membrane. Several models based on biophysical experiments have been proposed for their mechanism of action. Here we present the NMR-determined structure of the cyclic, cationic antimicrobial peptide cyclo(RRWWRF) both free in aqueous solution and bound to detergent micelles. The peptide has a rather flexible but ordered structure in water. A distinct structure is formed when the peptide is bound to a detergent micelle. The structures in neutral and negatively charged micelles are nearly identical but differ from that in aqueous solution. The orientation of the amino acid side chains creates an amphipathic molecule with the peptide backbone forming the hydrophilic part. The orientation of the peptide in the micelle was determined by using NOEs and paramagnetic agents. The peptide is oriented mainly parallel to the micelle surface in both detergents. Substitution of the arginine and tryptophan residues is known to influence the antimicrobial activity. Therefore the structure of the micelle-bound analogues cyclo(RRYYRF), cyclo(KKWWKF) and cyclo(RRNalNalRF) were also determined. They exhibit remarkable similarities in backbone conformation and side-chain orientation. The structure of these peptides allows the side-chain properties to be correlated to biological activity.
- Forschungsinstitut für Molekulare Pharmakologie, Robert-Rössle-Strasse 10, 13125 Berlin, Germany.
Organizational Affiliation: