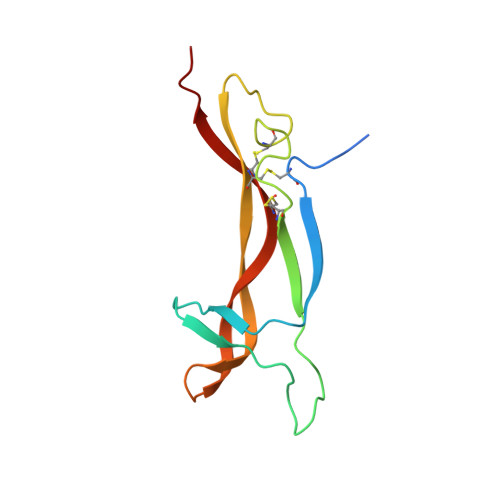
Structure of mouse 7S NGF: a complex of nerve growth factor with four binding proteins.
Bax, B., Blundell, T.L., Murray-Rust, J., McDonald, N.Q.(1997) Structure 5: 1275-1285
- PubMed: 9351801 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(97)00280-3
- Primary Citation Related Structures:
1SGF - PubMed Abstract:
Nerve growth factor (NGF) is a neurotrophic factor that promotes the differentiation and survival of certain populations of neurons in the central and peripheral nervous systems. 7S NGF is an alpha 2 beta 2 gamma 2 complex in which the beta-NGF dimer (the active neurotrophin) is associated with two alpha-NGF and two gamma-NGF subunits, which belong to the glandular kallikrein family of serine proteinases. The gamma-NGF subunit is an active serine proteinase capable of processing the precursor form of beta-NGF, whereas alpha-NGF is an inactive serine proteinase. The structure of 7S NGF could be used as a starting point to design inhibitors that prevent NGF binding to its receptors, as a potential treatment of neurodegenerative diseases. The crystal structure of 7S NGF shows that the two gamma-NGF subunits make extensive interactions with each other around the twofold axis of the complex and have the C-terminal residues of the beta-NGF subunits bound within their active sites. The 'activation domain' of each of the alpha-NGF subunits is in an inactive (zymogen-like) conformation and makes extensive interactions with the beta-NGF dimer. The two zinc ions that stabilize the complex are located at the relatively small interfaces between the alpha-NGF and gamma-NGF subunits. The structure of 7S NGF shows how the twofold axis of the central beta-NGF dimer organizes the symmetry of this multisubunit growth factor complex. The extensive surface of beta-NGF buried within the 7S complex explains the lack of neurotrophic activity observed for 7S NGF. The regions of the beta-NGF dimer that contact the alpha-NGF subunits overlap with those known to engage NGF receptors. Two disulphide-linked loops on alpha-NGF make multiple interactions with beta-NGF and suggest that it might be possible to design peptides that inhibit the binding of beta-NGF to its receptors.
- Department of Crystallography, Birkbeck College, London, UK. b.bax@cryst.bbk.ac.uk
Organizational Affiliation: