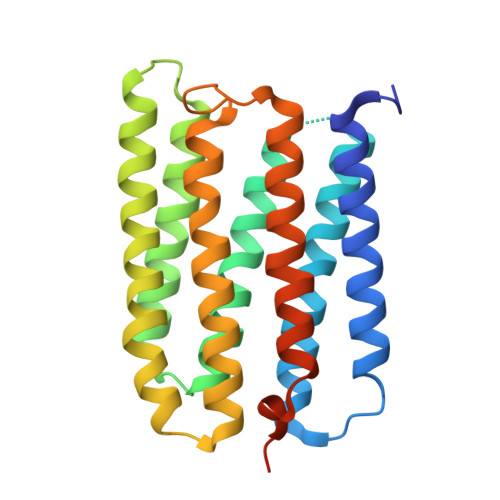
Specificity of anion binding in the substrate pocket of bacteriorhodopsin.
Facciotti, M.T., Cheung, V.S., Lunde, C.S., Rouhani, S., Baliga, N.S., Glaeser, R.M.(2004) Biochemistry 43: 4934-4943
- PubMed: 15109251 Search on PubMed
- DOI: https://doi.org/10.1021/bi035757s
- Primary Citation Related Structures:
1S8J, 1S8L - PubMed Abstract:
The structure of the D85S mutant of bacteriorhodopsin with a nitrate anion bound in the Schiff base binding site and the structure of the anion-free protein have been obtained in the same crystal form. Together with the previously solved structures of this anion pump, in both the anion-free state and bromide-bound state, these new structures provide insight into how this mutant of bacteriorhodopsin is able to bind a variety of different anions in the same binding pocket. The structural analysis reveals that the main structural change that accommodates different anions is the repositioning of the polar side chain of S85. On the basis of these X-ray crystal structures, the prediction is then made that the D85S/D212N double mutant might bind similar anions and do so over a broader pH range than does the single mutant. Experimental comparison of the dissociation constants, K(d), for a variety of anions confirms this prediction and demonstrates, in addition, that the binding affinity is dramatically improved by the D212N substitution.
- Life Sciences Division, Lawrence Berkeley National Laboratory, Berkeley, California 94720, USA.
Organizational Affiliation: