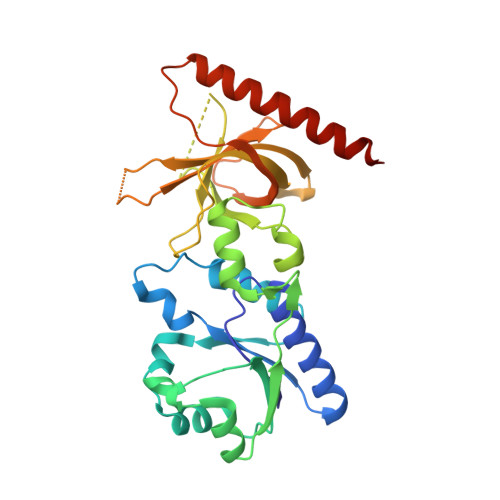
Crystal structure of flavin binding to FAD synthetase of Thermotoga maritima
Wang, W., Kim, R., Yokota, H., Kim, S.-H.(2005) Proteins 58: 246-248
- PubMed: 15468322 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20207
- Primary Citation Related Structures:
1S4M - Berkeley Structural Genomics Center, Physical Biosciences Division of the Lawrence Berkeley National Laboratory, Berkeley, California 94720, USA.
Organizational Affiliation: