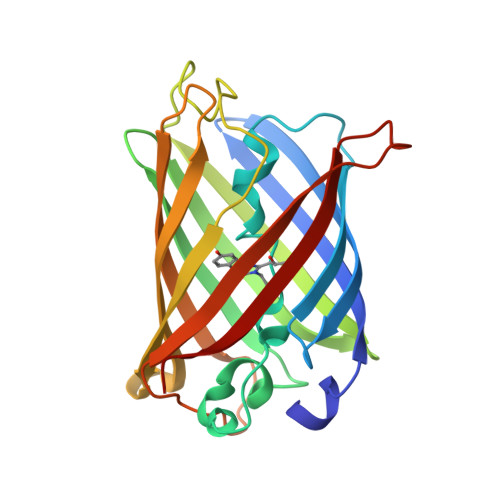
Probing the role of tryptophans in Aequorea victoria green fluorescent proteins with an expanded genetic code
Budisa, N., Pal, P.P., Alefelder, S., Birle, P., Krywcun, T., Rubini, M., Wenger, W., Bae, J.H., Steiner, T.(2004) Biol Chem 385: 191-202
- PubMed: 15101562 Search on PubMed
- DOI: https://doi.org/10.1515/BC.2004.038
- Primary Citation Related Structures:
1RM9, 1RMM, 1RMO, 1RMP - PubMed Abstract:
The expanded genetic code in combination with site-directed mutagenesis was used to probe spectroscopic and structural roles of tryptophan (Trp) residues in Aequorea victoria green fluorescent proteins (avGFPs). Nine different halogen-, chalcogen-, and methyl-containing Trp isosteric analogues and surrogates were incorporated into avGFPs containing indole moieties in, and outside of, the chromophore, by the use of the selective pressure incorporation method. Such isosteric replacements introduced minimal local geometry changes in indole moieties, often to the level of single atomic exchange ('atomic mutation') and do not affect three-dimensional structures of avGFPs but induce changes in spectral properties. Our approach offers a new platform to re-evaluate issues like resonance transfer, mechanisms of chromophore formation and maturation, as well as the importance of local geometry and weak sulphur-aromatic interactions for avGFP spectral properties and structural stability. The library of novel tailor-made avGFP mutants and variants generated in this work has demonstrated not only the potentials of the expanded genetic code to study spectroscopic functions, but also a new approach to generate tailor-made proteins with interesting and useful spectral properties.
- Max-Planck-Institut für Biochemie, Am Klopferspitz 18A, D-82152 Martinsried, Germany. budisa@biochem.mpg.de
Organizational Affiliation: