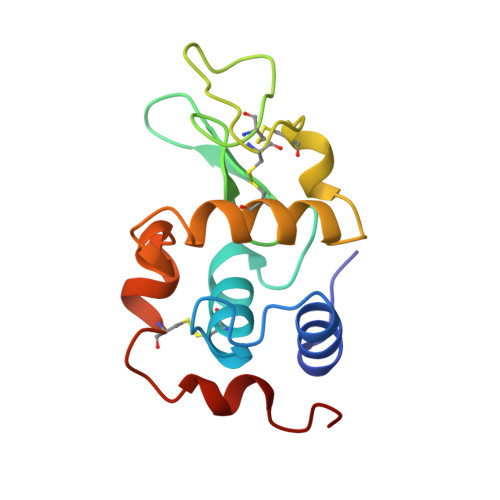
Crystal structure of a ubiquitin-dependent degradation substrate: a three-disulfide form of lysozyme.
Hill, C.P., Johnston, N.L., Cohen, R.E.(1993) Proc Natl Acad Sci U S A 90: 4136-4140
- PubMed: 8387211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.90.9.4136
- Primary Citation Related Structures:
1RCM - PubMed Abstract:
Covalent attachment of ubiquitin marks substrates for proteolysis, but features that identify ubiquitination targets such as chicken egg white lysozyme are poorly understood. Recognition of lysozyme first requires reduction of Cys-6 Cys-127, one of its four native disulfide bonds, and Cys-6,Cys-127-carboxymethylated (6,127-rcm) lysozyme can mimic this three-disulfide intermediate. The 6,127-rcm form of lysozyme is known to retain a substantially native-like conformation in solution, and we demonstrate that it is this folded structure that is recognized for ubiquitination. Because native lysozyme is not a substrate, differences between the native and three-disulfide structures must include features responsible for selective ubiquitination. The 1.9-A resolution crystal structure of 6,127-rcm-lysozyme, reported here, affords a view of this ubiquitin-dependent degradation substrate. Two conformers of 6,127-rcm-lysozyme were obtained in the crystal. These differ uniquely from crystal forms of native lysozyme by displacement of the C-terminal residues. The structures suggest that localized unfolding at the C terminus of three-disulfide lysozyme allows the complex of E3 alpha (ubiquitin-protein ligase) and E2 (ubiquitin-carrier protein) to bind to a surface that includes Lys-1 and the putative ubiquitination site Lys-13. From this we infer that the N-terminal and internal substrate recognition sites on the E3 alpha.E2 complex are separated by approximately 20 A.
- Department of Chemistry and Biochemistry, University of California, Los Angeles 90024-1570.
Organizational Affiliation: