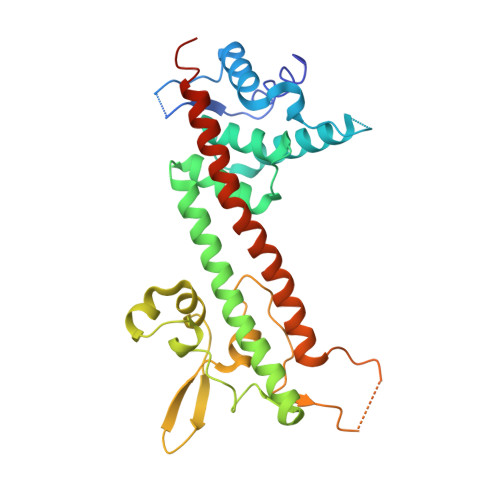
The structure of Yersinia pestis V-antigen, an essential virulence factor and mediator of immunity against plague
Derewenda, U., Mateja, A., Devedjiev, Y., Routzahn, K.M., Evdokimov, A.G., Derewenda, Z.S., Waugh, D.S.(2004) Structure 12: 301-306
- PubMed: 14962390 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.01.010
- Primary Citation Related Structures:
1R6F - PubMed Abstract:
The LcrV protein (V-antigen) is a multifunctional virulence factor in Yersinia pestis, the causative agent of plague. LcrV regulates the translocation of cytotoxic effector proteins from the bacterium into the cytosol of mammalian cells via a type III secretion system, possesses antihost activities of its own, and is also an active and passive mediator of resistance to disease. Although a crystal structure of this protein has been actively sought for better understanding of its role in pathogenesis, the wild-type LcrV was found to be recalcitrant to crystallization. We employed a surface entropy reduction mutagenesis strategy to obtain crystals of LcrV that diffract to 2.2 A and determined its structure. The refined model reveals a dumbbell-like molecule with a novel fold that includes an unexpected coiled-coil motif, and provides a detailed three-dimensional roadmap for exploring structure-function relationships in this essential virulence determinant.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, VA 22908, USA.
Organizational Affiliation: