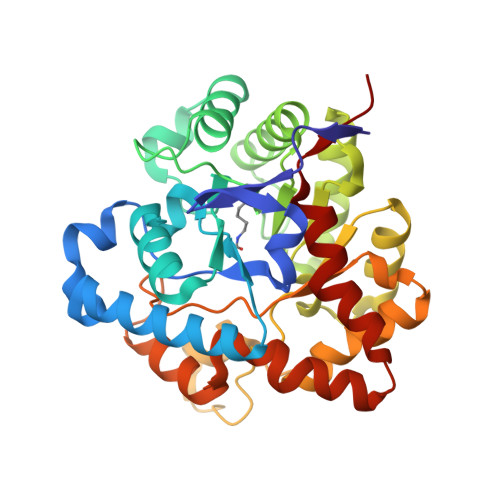
Structural and mutational studies of organophosphorus hydrolase reveal a cryptic and functional allosteric-binding site.
Grimsley, J.K., Calamini, B., Wild, J.R., Mesecar, A.D.(2005) Arch Biochem Biophys 442: 169-179
- PubMed: 16188223 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2005.08.012
- Primary Citation Related Structures:
1QW7 - PubMed Abstract:
Organophosphorus hydrolase detoxifies a broad range of organophosphate pesticides and the chemical warfare agents (CWAs) sarin and VX. Previously, rational genetic engineering produced OPH variants with 30-fold enhancements in the hydrolysis of CWA and their analogs. One interesting variant (H254R) in which the histidine at position 254 was changed to an arginine showed a 4-fold increase in the hydrolysis of demetonS (VX analog), a 14-fold decrease with paraoxon (an insecticide), and a 183-fold decrease with DFP (sarin analog). The three-dimensional structure of this enzyme at 1.9A resolution with the inhibitor, diethyl 4-methylbenzylphosphonate (EBP), revealed that the inhibitor did not bind at the active site, but bound exclusively into a well-defined surface pocket 12 A away from the active site. This structural feature was accompanied by non-competitive inhibition of paraoxon hydrolysis by EBP with H254R, in contrast to the native enzyme, which showed competitive inhibition. These parallel structure-function characteristics identify a functional, allosteric site on the surface of this enzyme.
- Biochemistry and Biophysics Department, Texas A&M University, College Station, TX 77843-2128, USA.
Organizational Affiliation: