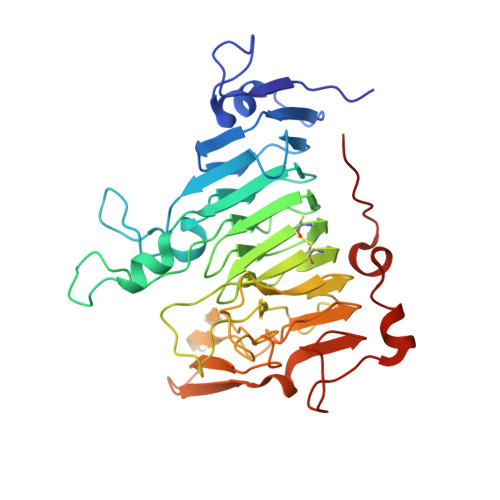
Three-Dimensional Structure of Erwinia Chrysanthemi Pectin Methylesterase Reveals a Novel Esterase Active Site
Jenkins, J., Mayans, O., Smith, D., Worboys, K., Pickersgill, R.(2001) J Mol Biology 305: 951
- PubMed: 11162105 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4324
- Primary Citation Related Structures:
1QJV - PubMed Abstract:
Most structures of neutral lipases and esterases have been found to adopt the common alpha/beta hydrolase fold and contain a catalytic Ser-His-Asp triad. Some variation occurs in both the overall protein fold and in the location of the catalytic triad, and in some enzymes the role of the aspartate residue is replaced by a main-chain carbonyl oxygen atom. Here, we report the crystal structure of pectin methylesterase that has neither the common alpha/beta hydrolase fold nor the common catalytic triad. The structure of the Erwinia chrysanthemi enzyme was solved by multiple isomorphous replacement and refined at 2.4 A to a conventional crystallographic R-factor of 17.9 % (R(free) 21.1 %). This is the first structure of a pectin methylesterase and reveals the enzyme to comprise a right-handed parallel beta-helix as seen in the pectinolytic enzymes pectate lyase, pectin lyase, polygalacturonase and rhamnogalacturonase, and unlike the alpha/beta hydrolase fold of rhamnogalacturonan acetylesterase with which it shares esterase activity. Pectin methylesterase has no significant sequence similarity with any protein of known structure. Sequence conservation among the pectin methylesterases has been mapped onto the structure and reveals that the active site comprises two aspartate residues and an arginine residue. These proposed catalytic residues, located on the solvent-accessible surface of the parallel beta-helix and in a cleft formed by external loops, are at a location similar to that of the active site and substrate-binding cleft of pectate lyase. The structure of pectin methylesterase is an example of a new family of esterases.
- Institute of Food Research, Norwich Research Park, Colney Lane, Norwich NR4 7UA, UK.
Organizational Affiliation: