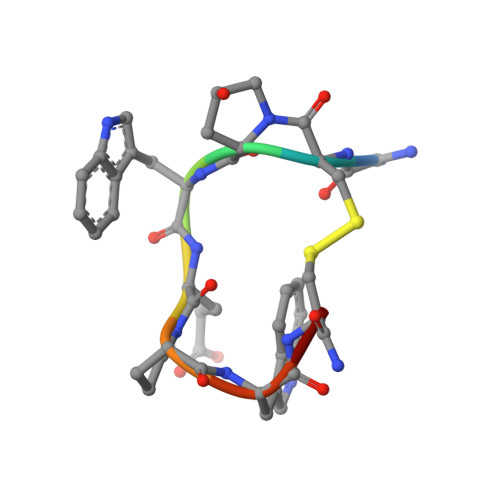
Solution structure of contryphan-R, a naturally occurring disulfide-bridged octapeptide containing D-tryptophan: comparison with protein loops.
Pallaghy, P.K., Melnikova, A.P., Jimenez, E.C., Olivera, B.M., Norton, R.S.(1999) Biochemistry 38: 11553-11559
- PubMed: 10471307
- DOI: https://doi.org/10.1021/bi990685j
- Primary Citation of Related Structures:
1QFB - PubMed Abstract:
Contryphan-R is a disulfide-constrained octapeptide containing a D-tryptophan that was isolated recently from venom of the cone shell Conus radiatus. The polypeptide is present in two forms in solution due to cis-trans isomerization at hydroxyproline 3. The solution structure of the major form of this unusual polypeptide, determined from NMR data, consists of a well-defined fold containing a non-hydrogen-bonded chain reversal from Gly1 to Glu5, which includes a cis-hydroxyproline and a D-Trp, and a type I beta-turn from Glu5 to Cys8. The presence of a putative salt bridge between the Glu5 carboxyl group and the N-terminal ammonium group is investigated by using various solvation models during energy minimization and is compared with the results of a pH titration. A comparison of the structure of contryphan-R with other cyclic peptide structures highlights some of the key structural determinants of these peptides and suggests that the contryphan-R fold could be exploited as a scaffold onto which unrelated protein binding surfaces could be grafted. Comparison with small disulfide-bridged loops in larger proteins shows that contryphan-R is similar to a commonly occurring loop structure found in proteins.
- Biomolecular Research Institute, Parkville, Australia.
Organizational Affiliation: